Selectivity changes during activation of mutant Shaker potassium channels
- PMID: 9236204
- PMCID: PMC2233792
- DOI: 10.1085/jgp.110.2.101
Selectivity changes during activation of mutant Shaker potassium channels
Abstract
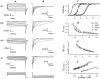
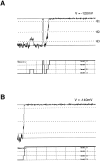
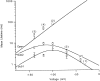
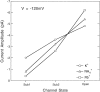
Mutations of the pore-region residue T442 in Shaker channels result in large effects on channel kinetics. We studied mutations at this position in the backgrounds of NH2-terminal-truncated Shaker H4 and a Shaker -NGK2 chimeric channel having high conductance (Lopez, G.A., Y.N. Jan, and L.Y. Jan. 1994. Nature (Lond.). 367: 179-182). While mutations of T442 to C, D, H, V, or Y resulted in undetectable expression in Xenopus oocytes, S and G mutants yielded functional channels having deactivation time constants and channel open times two to three orders of magnitude longer than those of the parental channel. Activation time courses at depolarized potentials were unaffected by the mutations, as were first-latency distributions in the T442S chimeric channel. The mutant channels show two subconductance levels, 37 and 70% of full conductance. From single-channel analysis, we concluded that channels always pass through the larger subconductance state on the way to and from the open state. The smaller subconductance state is traversed in approximately 40% of activation time courses. These states apparently represent kinetic intermediates in channel gating having voltage-dependent transitions with apparent charge movements of approximately 1.6 e0. The fully open T442S chimeric channel has the conductance sequence Rb+ > NH4+ > K+. The opposite conductance sequence, K+ > NH4+ > Rb+, is observed in each of the subconductance states, with the smaller subconductance state discriminating most strongly against Rb+.
Figures



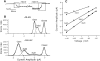




Similar articles
-
Intermediate conductances during deactivation of heteromultimeric Shaker potassium channels.J Gen Physiol. 1998 Oct;112(4):457-74. doi: 10.1085/jgp.112.4.457. J Gen Physiol. 1998. PMID: 9758864 Free PMC article.
-
Hidden Markov model analysis of intermediate gating steps associated with the pore gate of shaker potassium channels.J Gen Physiol. 2001 Nov;118(5):547-64. doi: 10.1085/jgp.118.5.547. J Gen Physiol. 2001. PMID: 11696611 Free PMC article.
-
Activation of Shaker potassium channels. III. An activation gating model for wild-type and V2 mutant channels.J Gen Physiol. 1998 Feb;111(2):313-42. doi: 10.1085/jgp.111.2.313. J Gen Physiol. 1998. PMID: 9450946 Free PMC article.
-
Binding of kappa-conotoxin PVIIA to Shaker K+ channels reveals different K+ and Rb+ occupancies within the ion channel pore.J Gen Physiol. 2004 Jul;124(1):71-81. doi: 10.1085/jgp.200409048. J Gen Physiol. 2004. PMID: 15226365 Free PMC article.
-
Emerging issues of connexin channels: biophysics fills the gap.Q Rev Biophys. 2001 Aug;34(3):325-472. doi: 10.1017/s0033583501003705. Q Rev Biophys. 2001. PMID: 11838236 Review.
Cited by
-
Localization of the activation gate of a voltage-gated Ca2+ channel.J Gen Physiol. 2005 Sep;126(3):205-12. doi: 10.1085/jgp.200509293. J Gen Physiol. 2005. PMID: 16129771 Free PMC article.
-
Saturation and microsecond gating of current indicate depletion-induced instability of the MaxiK selectivity filter.J Gen Physiol. 2007 Jul;130(1):83-97. doi: 10.1085/jgp.200709802. J Gen Physiol. 2007. PMID: 17591987 Free PMC article.
-
Two-dimensional kinetic analysis suggests nonsequential gating of mechanosensitive channels in Xenopus oocytes.Biophys J. 2001 Oct;81(4):2082-99. doi: 10.1016/S0006-3495(01)75857-2. Biophys J. 2001. PMID: 11566780 Free PMC article.
-
Control of voltage-gated K+ channel permeability to NMDG+ by a residue at the outer pore.J Gen Physiol. 2009 Apr;133(4):361-74. doi: 10.1085/jgp.200810139. J Gen Physiol. 2009. PMID: 19332619 Free PMC article.
-
A pain-inducing centipede toxin targets the heat activation machinery of nociceptor TRPV1.Nat Commun. 2015 Sep 30;6:8297. doi: 10.1038/ncomms9297. Nat Commun. 2015. PMID: 26420335 Free PMC article.
References
-
- Brown AM. Functional bases for interpreting amino acid sequences of voltage dependent K channels. Annu Rev Biophys Biomol Struct. 1993;22:173–198. - PubMed

