Mutational analysis of Mdm1p function in nuclear and mitochondrial inheritance
- PMID: 9245780
- PMCID: PMC2141631
- DOI: 10.1083/jcb.138.3.485
Mutational analysis of Mdm1p function in nuclear and mitochondrial inheritance
Abstract
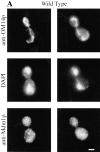
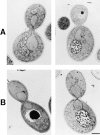
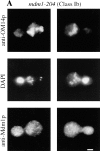
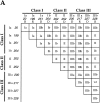
Nuclear and mitochondrial transmission to daughter buds of Saccharomyces cerevisiae depends on Mdm1p, an intermediate filament-like protein localized to numerous punctate structures distributed throughout the yeast cell cytoplasm. These structures disappear and organelle inheritance is disrupted when mdm1 mutant cells are incubated at the restrictive temperature. To characterize further the function of Mdm1p, new mutant mdm1 alleles that confer temperature-sensitive growth and defects in organelle inheritance but produce stable Mdm1p structures were isolated. Microscopic analysis of the new mdm1 mutants revealed three phenotypic classes: Class I mutants showed defects in both mitochondrial and nuclear transmission; Class II alleles displayed defective mitochondrial inheritance but had no effect on nuclear movement; and Class III mutants showed aberrant nuclear inheritance but normal mitochondrial distribution. Class I and II mutants also exhibited altered mitochondrial morphology, possessing primarily small, round mitochondria instead of the extended tubular structures found in wild-type cells. Mutant mdm1 alleles affecting nuclear transmission were of two types: Class Ia and IIIa mutants were deficient for nuclear movement into daughter buds, while Class Ib and IIIb mutants displayed a complete transfer of all nuclear DNA into buds. The mutations defining all three allelic classes mapped to two distinct domains within the Mdm1p protein. Genetic crosses of yeast strains containing different mdm1 alleles revealed complex genetic interactions including intragenic suppression, synthetic phenotypes, and intragenic complementation. These results support a model of Mdm1p function in which a network comprised of multimeric assemblies of the protein mediates two distinct cellular processes.
Figures





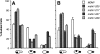

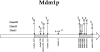

References
-
- Adams RJ, Pollard TD. Propulsion of organelles isolated from Acanthamoeba along actin filaments by myosin-I. Nature (Lond) 1986;322:754–756. - PubMed
-
- Allan V. Membrane traffic motors. FEBS (Fed Eur Biochem Soc) Lett. 1995;369:101–106. - PubMed
-
- Berger KH, Yaffe MP. Mitochondrial distribution and inheritance. Experientia (Basel) 1996;52:1111–1116. - PubMed
-
- Chen LB. Mitochondrial membrane potential in living cells. Annu Rev Cell Biol. 1988;4:155–181. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

