Quantitative single-cell-reverse transcription-PCR demonstrates that A-current magnitude varies as a linear function of shal gene expression in identified stomatogastric neurons
- PMID: 9254672
- PMCID: PMC6573138
- DOI: 10.1523/JNEUROSCI.17-17-06597.1997
Quantitative single-cell-reverse transcription-PCR demonstrates that A-current magnitude varies as a linear function of shal gene expression in identified stomatogastric neurons
Abstract
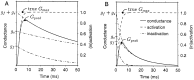
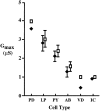
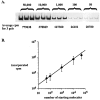
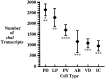
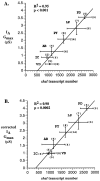
Different Shaker family alpha-subunit genes generate distinct voltage-dependent K+ currents when expressed in heterologous expression systems. Thus it generally is believed that diverse neuronal K+ current phenotypes arise, in part, from differences in Shaker family gene expression among neurons. It is difficult to evaluate the extent to which differential Shaker family gene expression contributes to endogenous K+ current diversity, because the specific Shaker family gene or genes responsible for a given K+ current are still unknown for nearly all adult neurons. In this paper we explore the role of differential Shaker family gene expression in creating transient K+ current (IA) diversity in the 14-neuron pyloric network of the spiny lobster, Panulirus interruptus. We used two-electrode voltage clamp to characterize the somatic IA in each of the six different cell types of the pyloric network. The size, voltage-dependent properties, and kinetic properties of the somatic IA vary significantly among pyloric neurons such that the somatic IA is unique in each pyloric cell type. Comparing these currents with the IAs obtained from oocytes injected with Panulirus shaker and shal cRNA (lobster Ishaker and lobster Ishal, respectively) reveals that the pyloric cell IAs more closely resemble lobster Ishal than lobster Ishaker. Using a novel, quantitative single-cell-reverse transcription-PCR method to count the number of shal transcripts in individual identified pyloric neurons, we found that the size of the somatic IA varies linearly with the number of endogenous shal transcripts. These data suggest that the shal gene contributes substantially to the peak somatic IA in all neurons of the pyloric network.
Figures



References
-
- Baker K, Salkoff L. The Drosophila shaker gene codes for a distinctive K+ current in a subset of neurons. Neuron. 1990;2:129–140. - PubMed
-
- Baro DJ, Cole CL, Harris-Warrick RM. RT-PCR analysis of shaker, shab, shaw, and shal gene expression in single neurons and glial cells. Receptors Channels. 1996b;4:149–159. - PubMed
-
- Barrantes FJ. Structural–functional correlates of the nicotinic acetylcholine receptor and its lipid microenvironment. FASEB J. 1993;7:1460–1467. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
