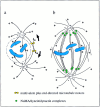
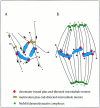
Pathways of spindle pole formation: different mechanisms; conserved components
- PMID: 9281574
- PMCID: PMC2136757
- DOI: 10.1083/jcb.138.5.953
Pathways of spindle pole formation: different mechanisms; conserved components
Figures
Similar articles
-
Spindle multipolarity is prevented by centrosomal clustering.Science. 2005 Jan 7;307(5706):127-9. doi: 10.1126/science.1104905. Science. 2005. PMID: 15637283
-
Only one spindle, if you please...Nat Cell Biol. 2006 Sep;8(9):901-2. doi: 10.1038/ncb0906-901. Nat Cell Biol. 2006. PMID: 16946735 No abstract available.
-
[Dynein and dynactin as organizers of the system of cell microtubules].Ontogenez. 2006 Sep-Oct;37(5):323-39. Ontogenez. 2006. PMID: 17066975 Review. Russian.
-
Examining how the spatial organization of chromatin signals influences metaphase spindle assembly.Nat Cell Biol. 2006 Sep;8(9):924-32. doi: 10.1038/ncb1455. Epub 2006 Aug 6. Nat Cell Biol. 2006. PMID: 16892054
-
Microtubule cytoskeleton: navigating the intracellular landscape.Curr Biol. 2003 May 27;13(11):R430-2. doi: 10.1016/s0960-9822(03)00362-2. Curr Biol. 2003. PMID: 12781150 Review.
Cited by
-
Mitotic progression and dual spindle formation caused by spindle association of de novo-formed microtubule-organizing centers in parthenogenetic embryos of Drosophila ananassae.Genetics. 2023 Feb 9;223(2):iyac178. doi: 10.1093/genetics/iyac178. Genetics. 2023. PMID: 36516293 Free PMC article.
-
Centrosome amplification is a frequent event in circulating tumor cells from subjects with metastatic breast cancer.Mol Oncol. 2020 Aug;14(8):1898-1909. doi: 10.1002/1878-0261.12687. Epub 2020 May 19. Mol Oncol. 2020. PMID: 32255253 Free PMC article.
-
Pericentrin and gamma-tubulin form a protein complex and are organized into a novel lattice at the centrosome.J Cell Biol. 1998 Apr 6;141(1):163-74. doi: 10.1083/jcb.141.1.163. J Cell Biol. 1998. PMID: 9531556 Free PMC article.
-
LIN-5 is a novel component of the spindle apparatus required for chromosome segregation and cleavage plane specification in Caenorhabditis elegans.J Cell Biol. 2000 Jan 10;148(1):73-86. doi: 10.1083/jcb.148.1.73. J Cell Biol. 2000. PMID: 10629219 Free PMC article.
-
MEI-1/MEI-2 katanin-like microtubule severing activity is required for Caenorhabditis elegans meiosis.Genes Dev. 2000 May 1;14(9):1072-84. Genes Dev. 2000. PMID: 10809666 Free PMC article.
References
-
- Afshar K, Barton NR, Hawley RS, Goldstein LSB. DNA binding and meiotic chromosomal localization of the Drosophilanod kinesin-like protein. Cell. 1995;81:129–138. - PubMed
-
- Boleti H, Karsenti E, Vernos I. Xklp2, a novel Xenopuscentrosomal kinesin-like protein required for centrosome separation during mitosis. Cell. 1996;84:49–59. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

