Enhancer complexes located downstream of both human immunoglobulin Calpha genes
- PMID: 9294139
- PMCID: PMC2199054
- DOI: 10.1084/jem.186.6.845
Enhancer complexes located downstream of both human immunoglobulin Calpha genes
Abstract
To investigate regulation of human immunoglobulin heavy chain expression, we have cloned DNA downstream from the two human Calpha genes, corresponding to the position in the mouse IgH cluster of a locus control region (LCR) that includes an enhancer which regulates isotype switching. Within 25 kb downstream of both the human immunoglobulin Calpha1 and Calpha2 genes we identified several segments of DNA which display B lymphoid-specific DNase I hypersensitivity as well as enhancer activity in transient transfections. The corresponding sequences downstream from each of the two human Calpha genes are nearly identical to each other. These enhancers are also homologous to three regions which lie in similar positions downstream from the murine Calpha gene and form the murine LCR. The strongest enhancers in both mouse and human have been designated HS12. Within a 135-bp core homology region, the human HS12 enhancers are approximately 90% identical to the murine homolog and include several motifs previously demonstrated to be important for function of the murine enhancer; additional segments of high sequence conservation suggest the possibility of previously unrecognized functional motifs. On the other hand, certain functional elements in the murine enhancer, including a B cell-specific activator protein site, do not appear to be conserved in human HS12. The human homologs of the murine enhancers designated HS3 and HS4 show lower overall sequence conservation, but for at least two of the functional motifs in the murine HS4 (a kappaB site and an octamer motif ) the human HS4 homologs are exactly conserved. An additional hypersensitivity site between human HS3 and HS12 in each human locus displays no enhancer activity on its own, but includes a region of high sequence conservation with mouse, suggesting the possibility of another novel functional element.
Figures


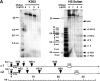







References
-
- Coffman RL, Lebman DA, Rothman P. The mechanism and regulation of immunoglobulin isotype switching. Adv Immunol. 1993;54:229–270. - PubMed
-
- Sideras P, Nilsson L, Islam KB, Quintana IZ, Freihof L, Rosen GJ, Hammarstrom L, Smith CI. Transcription of unrearranged Ig H chain genes in human B cell malignancies. Biased expression of genes encoded within the first duplication unit of the Ig H chain locus. J Immunol. 1992;149:244. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

