An RNA enhancer in a phage transcriptional antitermination complex functions as a structural switch
- PMID: 9303537
- PMCID: PMC275392
- DOI: 10.1101/gad.11.17.2214
An RNA enhancer in a phage transcriptional antitermination complex functions as a structural switch
Abstract
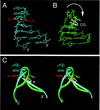
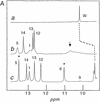
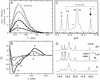
Antitermination protein N regulates the transcriptional program of phage lambda through recognition of RNA enhancer elements. Binding of an arginine-rich peptide to one face of an RNA hairpin organizes the other, which in turn binds to the host antitermination complex. The induced RNA structure mimics a GNRA hairpin, an organizational element of rRNA and ribozymes. The two faces of the RNA, bridged by a sheared GA base pair, exhibit a specific pattern of base stacking and base flipping. This pattern is extended by stacking of an aromatic amino acid side chain with an unpaired adenine at the N-binding surface. Such extended stacking is coupled to induction of a specific internal RNA architecture and is blocked by RNA mutations associated in vivo with loss of transcriptional antitermination activity. Mimicry of a motif of RNA assembly by an RNA-protein complex permits its engagement within the antitermination machinery.
Figures




References
-
- Albrechtsen B, Squires CL, Li S, Squires C. Antitermination of characterized transcriptional terminators by the Escherichia coli rrnG leader sequence. J Mol Biol. 1990;213:123–134. - PubMed
-
- Barik S, Ghosh B, Whalen W, Lazinski D, Das A. An antitermination protein engages the elongating transcription apparatus at a promoter-proximal recognition site. Cell. 1987;50:885–899. - PubMed
-
- Battiste JL, Tan R, Frankel AD, Williamson JR. Binding of an HIV Rev peptide to Rev responsive element RNA induces formation of purine-purine base pairs. Biochemistry. 1994;33:2741–2747. - PubMed
-
- Battiste JL, Mao H, Rao NS, Tan R, Muhandiram DR, Kay LE, Frankel AD, Williamson JR. Alpha helix-RNA major groove recognition in an HIV-1 rev peptide-RRE RNA complex. Science. 1996;273:1547–1551. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
