Mutations in dihydropteroate synthase are responsible for sulfone and sulfonamide resistance in Plasmodium falciparum
- PMID: 9391132
- PMCID: PMC28412
- DOI: 10.1073/pnas.94.25.13944
Mutations in dihydropteroate synthase are responsible for sulfone and sulfonamide resistance in Plasmodium falciparum
Abstract
Plasmodium falciparum causes the most severe form of malaria in humans. An important class of drugs in malaria treatment is the sulfone/sulfonamide group, of which sulfadoxine is the most commonly used. The target of sulfadoxine is the enzyme dihydropteroate synthase (DHPS), and sequencing of the DHPS gene has identified amino acid differences that may be involved in the mechanism of resistance to this drug. In this study we have sequenced the DHPS gene in 10 isolates from Thailand and identified a new allele of DHPS that has a previously unidentified amino acid difference. We have expressed eight alleles of P. falciparum PPPK-DHPS in Escherichia coli and purified the functional enzymes to homogeneity. Strikingly, the Ki for sulfadoxine varies by almost three orders of magnitude from 0.14 microM for the DHPS allele from sensitive isolates to 112 microM for an enzyme expressed in a highly resistant isolate. Comparison of the Ki of different sulfonamides and the sulfone dapsone has suggested that the amino acid differences in DHPS would confer cross-resistance to these compounds. These results show that the amino acid differences in the DHPS enzyme of sulfadoxine-resistant isolates of P. falciparum are central to the mechanism of resistance to sulfones and sulfonamides.
Figures
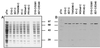
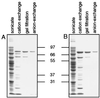

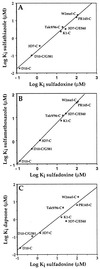
References
-
- Brown G M. Adv Enzymol Relat Areas Mol Biol. 1971;35:35–77. - PubMed
-
- Schellenberg K A, Coatney G R. Biochem Pharmacol. 1961;6:143–152. - PubMed
-
- Krungkrai J, Webster H K, Yuthavong Y. Mol Biochem Parasitol. 1989;32:25–38. - PubMed
-
- Ferone R. J Biol Chem. 1970;245:850–854. - PubMed
-
- Chulay J D, Watkins W M, Sixsmith D G. Am J Trop Med Hyg. 1984;33:325–330. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

