Regulation of transforming growth factor alpha expression in a growth factor-independent cell line
- PMID: 9418877
- PMCID: PMC121495
- DOI: 10.1128/MCB.18.1.303
Regulation of transforming growth factor alpha expression in a growth factor-independent cell line
Abstract
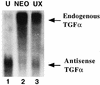
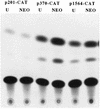
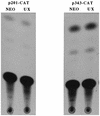
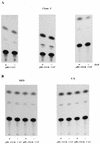
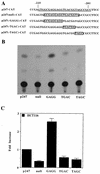
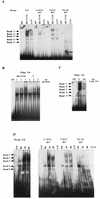
Aberrant transcriptional regulation of transforming growth factor alpha (TGF alpha) appears to be an important contributor to the malignant phenotype and the growth factor independence with which malignancy is frequently associated. However, little is known about the molecular mechanisms responsible for dysregulation of TGF alpha expression in the malignant phenotype. In this paper, we report on TGF alpha promoter regulation in the highly malignant growth factor-independent cell line HCT116. The HCT116 cell line expresses TGF alpha and the epidermal growth factor receptor (EGFR) but is not growth inhibited by antibodies to EGFR or TGF alpha. However, constitutive expression of TGF alpha antisense RNA in the HCT116 cell line resulted in the isolation of clones with markedly reduced TGF alpha mRNA and which were dependent on exogenous growth factors for proliferation. We hypothesized that if TGF alpha autocrine activation is the major stimulator of TGF alpha expression in this cell line, TGF alpha promoter activity should be reduced in the antisense TGF alpha clones in the absence of exogenous growth factor. This was the case. Moreover, transcriptional activation of the TGF alpha promoter was restored in an antisense-TGF alpha-mRNA-expressing clone which had reverted to a growth factor-independent phenotype. Using this model system, we were able to identify a 25-bp element within the TGF alpha promoter which conferred TGF alpha autoregulation to the TGF alpha promoter in the HCT116 cell line. In the TGF alpha-antisense-RNA-expressing clones, this element was activated by exogenous EGF. This 25-bp sequence contained no consensus sequences of known transcription factors so that the TGF alpha or EGF regulatory element within this 25-bp sequence represents a unique element. Further characterization of this 25-bp DNA sequence by deletion analysis revealed that regulation of TGF alpha promoter activity by this sequence is complex, as both repressors and activators bind in this region, but the overall expression of the activators is pivotal in determining the level of response to EGF or TGF alpha stimulation. The specific nuclear proteins binding to this region are also regulated in an autocrine-TGF alpha-dependent fashion and by exogenous EGF in EGF-deprived TGF alpha antisense clone 33. This regulation is identical to that seen in the growth factor-dependent cell line FET, which requires exogenous EGF for optimal growth. Moreover, the time response of the stimulation of trans-acting factor binding by EGF suggests that the effect is directly due to growth factor and not mediated by changes in growth state. We conclude that this element appears to represent the major positive regulator of TGF alpha expression in the growth factor-independent HCT116 cell line and may represent the major site of transcriptional dysregulation of TGF alpha promoter activity in the growth factor-independent phenotype.
Figures


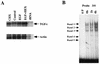
Similar articles
-
Post-transcriptional control regulates transforming growth factor alpha in the human carcinoma KB cell line.J Biol Chem. 1996 Nov 22;271(47):30290-6. doi: 10.1074/jbc.271.47.30290. J Biol Chem. 1996. PMID: 8939983
-
The role of transforming growth factor alpha in determining growth factor independence.Cancer Res. 2003 Aug 1;63(15):4731-8. Cancer Res. 2003. PMID: 12907656
-
The autocrine loop of epidermal growth factor receptor-epidermal growth factor / transforming growth factor-alpha in malignant rhabdoid tumor cell lines: heterogeneity of autocrine mechanism in TTC549.Jpn J Cancer Res. 2001 Mar;92(3):269-78. doi: 10.1111/j.1349-7006.2001.tb01091.x. Jpn J Cancer Res. 2001. PMID: 11267936 Free PMC article.
-
Growth factors in progression of human esophageal and gastric carcinomas.Exp Pathol. 1990;40(4):291-300. doi: 10.1016/s0232-1513(11)80316-6. Exp Pathol. 1990. PMID: 2098274 Review.
-
Transforming growth factor alpha: Key insights into physiological role, cancer therapeutics, and biomarker potential (A review).Int J Biol Macromol. 2025 May;310(Pt 2):143212. doi: 10.1016/j.ijbiomac.2025.143212. Epub 2025 Apr 17. Int J Biol Macromol. 2025. PMID: 40250676 Review.
Cited by
-
Role of protein kinase C and epidermal growth factor receptor signalling in growth stimulation by neurotensin in colon carcinoma cells.BMC Cancer. 2011 Oct 2;11:421. doi: 10.1186/1471-2407-11-421. BMC Cancer. 2011. PMID: 21961726 Free PMC article.
-
P53-mediated induction of Cox-2 counteracts p53- or genotoxic stress-induced apoptosis.EMBO J. 2002 Nov 1;21(21):5635-44. doi: 10.1093/emboj/cdf591. EMBO J. 2002. PMID: 12411481 Free PMC article.
-
Pancreatic cancer cells require an EGF receptor-mediated autocrine pathway for proliferation in serum-free conditions.Br J Cancer. 2001 Apr 6;84(7):926-35. doi: 10.1054/bjoc.2001.1698. Br J Cancer. 2001. PMID: 11286473 Free PMC article.
-
Formaldehyde induces micronuclei in mouse erythropoietic cells and suppresses the expansion of human erythroid progenitor cells.Toxicol Lett. 2014 Jan 13;224(2):233-9. doi: 10.1016/j.toxlet.2013.10.028. Epub 2013 Nov 1. Toxicol Lett. 2014. PMID: 24188930 Free PMC article.
References
-
- Angel P, Imagawa M, Chiu R, Stein B, Imbra R J, Rahmsdorf H J, Jonat C, Herrlich P, Karin M. Phorbol ester-inducible genes contain a common cis element recognized by a TPA-modulated trans-acting factor. Cell. 1987;49:729–739. - PubMed
-
- Bjorge J D, Paterson A J, Kudlow J E. Phorbol ester or epidermal growth factor (EGF) stimulates the concurrent accumulation of mRNA for the EGF receptor and its ligand transforming growth factor-α in a breast cancer cell line. J Biol Chem. 1989;264:4021–4027. - PubMed
-
- Boyd D D, Levine A E, Brattain D E, McKnight M K, Brattain M G. A comparison of growth requirements for two human intratumoral colon carcinoma cell lines in monolayer and soft agarose. Cancer Res. 1988;48:2469–2474. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
