Mutations in the human lambda5/14.1 gene result in B cell deficiency and agammaglobulinemia
- PMID: 9419212
- PMCID: PMC2199185
- DOI: 10.1084/jem.187.1.71
Mutations in the human lambda5/14.1 gene result in B cell deficiency and agammaglobulinemia
Abstract
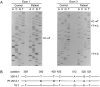
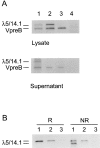
B cell precursors transiently express a pre-B cell receptor complex consisting of a rearranged mu heavy chain, a surrogate light chain composed of lambda5/14.1 and VpreB, and the immunoglobulin (Ig)-associated signal transducing chains, Igalpha and Igbeta. Mutations in the mu heavy chain are associated with a complete failure of B cell development in both humans and mice, whereas mutations in murine lambda5 result in a leaky phenotype with detectable humoral responses. In evaluating patients with agammaglobulinemia and markedly reduced numbers of B cells, we identified a boy with mutations on both alleles of the gene for lambda5/14.1. The maternal allele carried a premature stop codon in the first exon of lambda5/14.1 and the paternal allele demonstrated three basepair substitutions in a 33-basepair sequence in exon 3. The three substitutions correspond to the sequence in the lambda5/14. 1 pseudogene 16.1 and result in an amino acid substitution at an invariant proline. When expressed in COS cells, the allele carrying the pseudogene sequence resulted in defective folding and secretion of mutant lambda5/14.1. These findings indicate that expression of the functional lambda5/14.1 is critical for B cell development in the human.
Figures



References
-
- Spanopoulou E, Roman CA, Corcoran LM, Schlissel MS, Silver DP, Nemazee D, Nussenzweig MC, Shinton SA, Hardy RR, Baltimore D. Functional immunoglobulin transgenes guide ordered B-cell differentiation in Rag-1–deficient mice. Genes Dev. 1994;8:1030–1042. - PubMed
-
- Young F, Ardman B, Shinkai Y, Lansford R, Blackwell TK, Mendelsohn M, Rolink A, Melchers F, Alt FW. Influence of immunoglobulin heavy- and light-chain expression on B-cell differentiation. Genes Dev. 1994;8:1043–1057. - PubMed
-
- Rajewsky K. Clonal selection and learning in the antibody system. Nature. 1996;381:751–758. - PubMed
-
- Burrows PD, Cooper MD. B cell development and differentiation. Curr Opin Immunol. 1997;9:239–244. - PubMed
-
- Yel L, Minegishi Y, Coustan-Smith E, Buckley RH, Trubel H, Pachman LM, Kitchingman GR, Campana D, Rohrer J, Conley MD. Mutations in the mu heavy chain gene in patients with agammaglobulinemia. N Engl J Med. 1996;335:1486–1493. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

