The variable domain of nonassembled Ig light chains determines both their half-life and binding to the chaperone BiP
- PMID: 9465057
- PMCID: PMC19100
- DOI: 10.1073/pnas.95.4.1574
The variable domain of nonassembled Ig light chains determines both their half-life and binding to the chaperone BiP
Abstract
Not much is known about the features that determine the biological stability of a molecule retained in the endoplasmic reticulum (ER). Ig light (L) chains that are not secreted in the absence of Ig heavy (H) chain expression bind to the ER chaperone BiP as partially folded molecules until they are degraded. Although all Ig L chains have the same three-dimensional structure when part of an antibody molecule, the degradation rate of unassembled Ig L chains is not identical. For instance, the two nonsecreted murine Ig L chains, kappaNS1 and lambdaFS62, are degraded with half-lives of approximately 1 and 4 hr, respectively, in the same NS1 myeloma cells. Furthermore, the BiP/lambdaFS62 Ig L chain complex appears to be more stable than the BiP/kappaNS1 complex. Here, we used the ability of single Ig domains to form an internal disulfide bond after folding as a measure of the folding state of kappaNS1 and lambdaFS62 Ig L chains. Both of these nonsecreted L chains lack the internal disulfide bond in the variable (V) domain, whereas the constant (C) domain was folded in that respect. In both cases the unfolded V domain provided the BiP binding site. The stability of BiP binding to these two nonsecreted proteins was quite different, and both the stability of the BiP:Ig L chain complex and the half-life of the Ig L chain could be transferred from one Ig L chain isotype to the other by swapping the V domains. Our data suggest that the physical stability of BiP association with an unfolded region of a given light chain determines the half-life of that light chain, indicating a direct link between chaperone interaction and delivery of partially folded substrates to the mammalian degradation machinery.
Figures
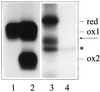


Similar articles
-
Molecular chaperones involved in protein degradation in the endoplasmic reticulum: quantitative interaction of the heat shock cognate protein BiP with partially folded immunoglobulin light chains that are degraded in the endoplasmic reticulum.Proc Natl Acad Sci U S A. 1995 Feb 28;92(5):1764-8. doi: 10.1073/pnas.92.5.1764. Proc Natl Acad Sci U S A. 1995. PMID: 7878056 Free PMC article.
-
BiP and immunoglobulin light chain cooperate to control the folding of heavy chain and ensure the fidelity of immunoglobulin assembly.Mol Biol Cell. 1999 Jul;10(7):2209-19. doi: 10.1091/mbc.10.7.2209. Mol Biol Cell. 1999. PMID: 10397760 Free PMC article.
-
Mapping the major interaction between binding protein and Ig light chains to sites within the variable domain.J Immunol. 1999 Oct 1;163(7):3842-50. J Immunol. 1999. PMID: 10490983
-
Biosynthesis, assembly and secretion of coagulation factor VIII.Blood Coagul Fibrinolysis. 1997 Dec;8 Suppl 2:S3-14. Blood Coagul Fibrinolysis. 1997. PMID: 9607108 Review.
-
BiP binding keeps ATF6 at bay.Dev Cell. 2002 Jul;3(1):1-2. doi: 10.1016/s1534-5807(02)00210-1. Dev Cell. 2002. PMID: 12110159 Review.
Cited by
-
ERdj3, a stress-inducible endoplasmic reticulum DnaJ homologue, serves as a cofactor for BiP's interactions with unfolded substrates.Mol Biol Cell. 2005 Jan;16(1):40-50. doi: 10.1091/mbc.e04-05-0434. Epub 2004 Nov 3. Mol Biol Cell. 2005. PMID: 15525676 Free PMC article.
-
Dissection of structural and functional requirements that underlie the interaction of ERdj3 protein with substrates in the endoplasmic reticulum.J Biol Chem. 2014 Oct 3;289(40):27504-12. doi: 10.1074/jbc.M114.587147. Epub 2014 Aug 20. J Biol Chem. 2014. PMID: 25143379 Free PMC article.
-
The Endoplasmic Reticulum-associated Degradation of Transthyretin Variants Is Negatively Regulated by BiP in Mammalian Cells.J Biol Chem. 2009 Mar 27;284(13):8312-21. doi: 10.1074/jbc.M809354200. Epub 2009 Feb 2. J Biol Chem. 2009. PMID: 19188365 Free PMC article.
-
Protein Quality Control in the Endoplasmic Reticulum.Protein J. 2019 Jun;38(3):317-329. doi: 10.1007/s10930-019-09831-w. Protein J. 2019. PMID: 31004255 Free PMC article. Review.
-
Characterization of an ERAD pathway for nonglycosylated BiP substrates, which require Herp.Mol Cell. 2007 Nov 30;28(4):544-54. doi: 10.1016/j.molcel.2007.09.012. Mol Cell. 2007. PMID: 18042451 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

