Multiple functions within the Epstein-Barr virus EBNA-3A protein
- PMID: 9499037
- PMCID: PMC109476
- DOI: 10.1128/JVI.72.3.1862-1869.1998
Multiple functions within the Epstein-Barr virus EBNA-3A protein
Abstract
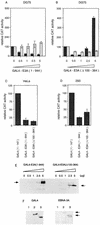
Two regions of the EBNA-3A protein of Epstein-Barr virus were shown to be capable of binding to the cell protein RBP-Jk (also known as CBF-1), a component of the Notch signaling pathway. Consistent with this binding, EBNA-3A inhibited reporter gene expression from plasmids containing RBP-Jk DNA binding sites within their promoters, including the Cp promoter. When EBNA-3A was linked to a GAL4 DNA binding domain, it repressed the activity of a promoter containing GAL4 binding sites at all plasmid concentrations tested. However, a deletion mutant of EBNA-3A lacking amino acids 100 to 364 showed a biphasic response in the GAL4 assay: it inhibited transcription at low DNA concentrations but activated it at high DNA concentrations. There appears to be a gene activation function within EBNA-3A that is masked in the full-length protein in this assay. Current models for EBNA-3 function have stressed transcription repression through binding to RBP-Jk, but we consider an alternative scheme in which the role of the binding of EBNA-3A, -3B, and -3C to RBP-Jk is to buffer the levels of active EBNA-3 protein. We have also found that the behavior of EBNA-3A in a cell fractionation procedure that distinguishes insoluble matrix from soluble cell fractions is modified by EBNA-LP, indicating a further novel level of interplay between the EBNA proteins.
Figures

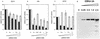

References
-
- Allan G J, Inman G J, Parker B D, Rowe D T, Farrell P J. Cell growth effects of Epstein-Barr virus leader protein. J Gen Virol. 1992;73:1547–1551. - PubMed
-
- Baer R, Bankier A T, Biggin M D, Deininger P L, Farrell P J, Gibson T J, Hatfull G, Hudson G S, Satchwell S C, Seguin C, et al. DNA sequence and expression of the B95-8 Epstein-Barr virus genome. Nature. 1984;310:207–211. - PubMed
-
- Ben-Bassat H, Goldblum N, Mitrani S, Goldblum T, Yoffey J M, Cohen M M, Bentwich Z, Ramot B, Klein E, Klein G. Establishment in continuous culture of a new type of lymphocyte from a “Burkitt like” malignant lymphoma (line D.G.-75) Int J Cancer. 1977;19:27–33. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

