Sequence-specific binding of human immunodeficiency virus type 1 nucleocapsid protein to short oligonucleotides
- PMID: 9499042
- PMCID: PMC109481
- DOI: 10.1128/JVI.72.3.1902-1909.1998
Sequence-specific binding of human immunodeficiency virus type 1 nucleocapsid protein to short oligonucleotides
Abstract
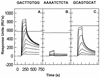
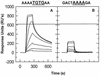
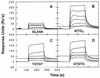
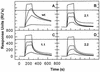
We have analyzed the binding of recombinant human immunodeficiency virus type 1 nucleocapsid protein (NC) to very short oligonucleotides by using surface plasmon resonance (SPR) technology. Our experiments, which were conducted at a moderate salt concentration (0.15 M NaCl), showed that NC binds more stably to runs of d(G) than to other DNA homopolymers. However, it exhibits far more stable binding with the alternating base sequence d(TG)n than with any homopolymeric oligodeoxyribonucleotide; thus, it shows a strong sequence preference under our experimental conditions. We found that the minimum length of an alternating d(TG) sequence required for stable binding was five nucleotides. Stable binding to the tetranucleotide d(TG)2 was observed only under conditions where two tetranucleotide molecules were held in close spatial proximity. The stable, sequence-specific binding to d(TG)n required that both zinc fingers be present, each in its proper position in the NC protein, and was quite salt resistant, indicating a large hydrophobic contribution to the binding. Limited tests with RNA oligonucleotides indicated that the preferential sequence-specific binding observed with DNA also occurs with RNA. Evidence was also obtained that NC can bind to nucleic acid molecules in at least two distinct modes. The biological significance of the specific binding we have detected is not known; it may reflect the specificity with which the parent Gag polyprotein packages genomic RNA or may relate to the functions of NC after cleavage of the polyprotein, including its role as a nucleic acid chaperone.
Figures




Similar articles
-
Nucleocapsid and matrix protein contributions to selective human immunodeficiency virus type 1 genomic RNA packaging.J Virol. 1998 Mar;72(3):1983-93. doi: 10.1128/JVI.72.3.1983-1993.1998. J Virol. 1998. PMID: 9499052 Free PMC article.
-
In vitro selection of RNAs that bind to the human immunodeficiency virus type-1 gag polyprotein.Nucleic Acids Res. 1997 Jul 15;25(14):2902-10. doi: 10.1093/nar/25.14.2902. Nucleic Acids Res. 1997. PMID: 9207041 Free PMC article.
-
HIV-1 nucleocapsid protein as a nucleic acid chaperone: spectroscopic study of its helix-destabilizing properties, structural binding specificity, and annealing activity.J Mol Biol. 2002 May 3;318(3):749-64. doi: 10.1016/S0022-2836(02)00043-8. J Mol Biol. 2002. PMID: 12054820
-
Nucleic acid sequence discrimination by the HIV-1 nucleocapsid protein NCp7: a fluorescence study.Biochemistry. 1999 Dec 21;38(51):16816-25. doi: 10.1021/bi991145p. Biochemistry. 1999. PMID: 10606514
-
HIV-1 nucleocapsid protein activates transient melting of least stable parts of the secondary structure of TAR and its complementary sequence.J Mol Biol. 2002 Mar 29;317(3):385-99. doi: 10.1006/jmbi.2002.5429. J Mol Biol. 2002. PMID: 11922672
Cited by
-
Overview of the Nucleic-Acid Binding Properties of the HIV-1 Nucleocapsid Protein in Its Different Maturation States.Viruses. 2020 Sep 29;12(10):1109. doi: 10.3390/v12101109. Viruses. 2020. PMID: 33003650 Free PMC article. Review.
-
Insights on the role of nucleic acid/protein interactions in chaperoned nucleic acid rearrangements of HIV-1 reverse transcription.Proc Natl Acad Sci U S A. 2007 Mar 27;104(13):5261-7. doi: 10.1073/pnas.0700166104. Epub 2007 Mar 19. Proc Natl Acad Sci U S A. 2007. PMID: 17372205 Free PMC article.
-
Characterization of Rous sarcoma virus Gag particles assembled in vitro.J Virol. 2001 Mar;75(6):2753-64. doi: 10.1128/JVI.75.6.2753-2764.2001. J Virol. 2001. PMID: 11222698 Free PMC article.
-
Mechanism for nucleic acid chaperone activity of HIV-1 nucleocapsid protein revealed by single molecule stretching.Proc Natl Acad Sci U S A. 2001 May 22;98(11):6121-6. doi: 10.1073/pnas.101033198. Epub 2001 May 8. Proc Natl Acad Sci U S A. 2001. PMID: 11344257 Free PMC article.
-
Nucleocapsid promotes localization of HIV-1 gag to uropods that participate in virological synapses between T cells.PLoS Pathog. 2010 Oct 28;6(10):e1001167. doi: 10.1371/journal.ppat.1001167. PLoS Pathog. 2010. PMID: 21060818 Free PMC article.
References
-
- Allen P, Collins B, Brown D, Hostomsky Z, Gold L. A specific RNA structural motif mediates high affinity binding by the HIV-1 nucleocapsid protein (NCp7) Virology. 1996;225:306–315. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

