E2F-6, a member of the E2F family that can behave as a transcriptional repressor
- PMID: 9501179
- PMCID: PMC19658
- DOI: 10.1073/pnas.95.6.2850
E2F-6, a member of the E2F family that can behave as a transcriptional repressor
Abstract
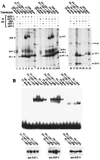
The E2F family of proteins is required to establish the correct cell-cycle-dependent transcription of genes that direct the process of cell division. All previously identified E2F proteins can act in a similar manner; depending on whether or not they are associated with the cell cycle inhibitors the retinoblastoma protein (pRB), p107, or p130, they can either repress or activate the transcription of E2F-responsive genes. We now report the cloning and characterization of another E2F family member, E2F-6, whose structure is reminiscent of the dominant inhibitors of other transcription factor families. The dimerization and DNA binding properties of E2F-6 are similar to those of the other E2F family members. However, it is not regulated by pRB, p107, or p130, and it is unable to activate transcription. Instead, it can act to repress the transcription of E2F responsive genes by countering the activity of the other E2F complexes via a pRB-, p107-, or p130-independent mechanism.
Figures


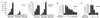
References
-
- Weinberg R A. Cancer Surv. 1992;12:43–57. - PubMed
-
- Dyson N, Harlow E. Cancer Surv. 1992;12:161–195. - PubMed
-
- Bartek J, Bartkova J, Lukas J. Curr Opin Cell Biol. 1996;8:805–814. - PubMed
-
- Pollock P M, Pearson J V, Hayward N K. Genes Chromosomes Cancer. 1996;15:77–88. - PubMed
-
- Nevins J R. Nature (London) 1992;358:375–376. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

