Clinical comparison of an enhanced-sensitivity branched-DNA assay and reverse transcription-PCR for quantitation of human immunodeficiency virus type 1 RNA in plasma
- PMID: 9508301
- PMCID: PMC104614
- DOI: 10.1128/JCM.36.3.716-720.1998
Clinical comparison of an enhanced-sensitivity branched-DNA assay and reverse transcription-PCR for quantitation of human immunodeficiency virus type 1 RNA in plasma
Abstract
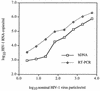
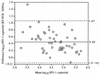
The performance characteristics of an enhanced-sensitivity branched-DNA assay (bDNA) (Quantiplex HIV-1 version 2.0; Chiron Corp., Emeryville, Calif.) and a reverse transcription (RT)-PCR assay (AMPLICOR HIV-1 Monitor; Roche Diagnostic Systems, Inc., Branchburg, N.J.) were compared in a molecular diagnostic laboratory. Samples used in this evaluation included linearity and reproducibility panels made by dilution of a human immunodeficiency virus type 1 (HIV-1) stock culture of known virus particle count in HIV-1-negative plasma, a subtype panel consisting of HIV-1 subtypes A through F at a standardized level, and 64 baseline plasma specimens from HIV-1-infected individuals. Plots of log10 HIV RNA copies per milliliter versus log10 nominal virus particles per milliliter demonstrated that both assays were linear over the stated dynamic ranges (bDNA, r = 0.98; RT-PCR, r = 0.99), but comparison of the slopes of the regression lines (bDNA, m = 0.96; RT-PCR, m = 0.83) suggested that RT-PCR had greater proportional systematic error. The between-run coefficients of variation for bDNA and RT-PCR were 24.3 and 34.3%, respectively, for a sample containing 1,650 nominal virus particles/ml and 44.0 and 42.7%, respectively, for a sample containing 165 nominal virus particles/ml. Subtypes B, C, and D were quantitated with similar efficiencies by bDNA and RT-PCR; however, RT-PCR was less efficient in quantitating subtypes A, E, and F. One non-B subtype was recognized in our clinical specimens based on the ratio of values obtained with the two methods. HIV-1 RNA was quantitated in 53 (83%) baseline plasma specimens by bDNA and in 55 (86%) specimens by RT-PCR. RT-PCR values were consistently greater than bDNA values, with population means of 142,419 and 67,580 copies/ml, respectively (P < 0.01). The results were highly correlated (r = 0.91), but the agreement was poor (mean difference in log10 copies per milliliter +/- 2 standard deviations, 0.45 +/- 0.61) for the 50 clinical specimens that gave discrete values with both methods.
Figures

References
-
- Bland J M, Altman D G. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet. 1986;i:307–310. - PubMed
-
- Caliendo A M. Methods, interpretation and application of HIV-1 viral load measurements. Clin Microbiol Newslett. 1997;19:1–5.
-
- Clewley J P. Genetic diversity and HIV detection by PCR. Lancet. 1995;346:1489. . (Letter.) - PubMed
-
- Costa J, Montes B, Reynes J, Peeters M, Segarra C, Vendrell J P, Delaporte E, Segondy M. Comparative evaluation of three assays for the quantitation of human immunodeficiency virus type 1 RNA in plasma. J Med Virol. 1996;50:293–302. - PubMed
-
- Dunne A L, Crowe S M. Comparison of branched DNA and reverse transcriptase polymerase chain reaction for quantifying six different HIV-1 subtypes in plasma. AIDS. 1997;11:2–3. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

