Partial purification and characterization of the maize mitochondrial pyruvate dehydrogenase complex
- PMID: 9536062
- PMCID: PMC35052
- DOI: 10.1104/pp.116.4.1443
Partial purification and characterization of the maize mitochondrial pyruvate dehydrogenase complex
Abstract
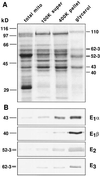
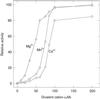
The pyruvate dehydrogenase complex was partially purified and characterized from etiolated maize (Zea mays L.) shoot mitochondria. Analysis by sodium dodecyl sulfate-polyacrylamide gel electrophoresis showed proteins of 40, 43, 52 to 53, and 62 to 63 kD. Immunoblot analyses identified these proteins as the E1beta-, E1alpha-, E2-, and E3-subunits, respectively. The molecular mass of maize E2 is considerably smaller than that of other plant E2 subunits (76 kD). The activity of the maize mitochondrial complex has a pH optimum of 7.5 and a divalent cation requirement best satisfied by Mg2+. Michaelis constants for the substrates were 47, 3, 77, and 1 &mgr;m for pyruvate, coenzyme A (CoA), NAD+, and thiamine pyrophosphate, respectively. The products NADH and acetyl-CoA were competitive inhibitors with respect to NAD+ and CoA, and the inhibition constants were 15 and 47 &mgr;m, respectively. The complex was inactivated by phosphorylation and was reactivated after the removal of ATP and the addition of Mg2+.
Figures



Similar articles
-
Plant mitochondrial pyruvate dehydrogenase complex: purification and identification of catalytic components in potato.Biochem J. 1998 Sep 15;334 ( Pt 3)(Pt 3):571-6. doi: 10.1042/bj3340571. Biochem J. 1998. PMID: 9729464 Free PMC article.
-
Some kinetic and regulatory properties of the pea mitochondrial pyruvate dehydrogenase complex.Plant Physiol. 1987 Feb;83(2):306-10. doi: 10.1104/pp.83.2.306. Plant Physiol. 1987. PMID: 16665241 Free PMC article.
-
Plant pyruvate dehydrogenase complex purification, characterization and regulation by metabolites and phosphorylation.Biochim Biophys Acta. 1977 Dec 8;485(2):336-49. doi: 10.1016/0005-2744(77)90169-3. Biochim Biophys Acta. 1977. PMID: 922017
-
The elementary reactions of the pig heart pyruvate dehydrogenase complex. A study of the inhibition by phosphorylation.Biochem J. 1976 Jul 1;157(1):41-67. doi: 10.1042/bj1570041. Biochem J. 1976. PMID: 183746 Free PMC article.
-
Purification and Characterization of the Pea Chloroplast Pyruvate Dehydrogenase Complex : A Source of Acetyl-CoA and NADH for Fatty Acid Biosynthesis.Plant Physiol. 1985 Mar;77(3):571-7. doi: 10.1104/pp.77.3.571. Plant Physiol. 1985. PMID: 16664100 Free PMC article.
Cited by
-
Temperature Shift Experiments Suggest That Metabolic Impairment and Enhanced Rates of Photorespiration Decrease Organic Acid Levels in Soybean Leaflets Exposed to Supra-Optimal Growth Temperatures.Metabolites. 2015 Aug 5;5(3):443-54. doi: 10.3390/metabo5030443. Metabolites. 2015. PMID: 26251925 Free PMC article.
-
Biochemistry, proteomics, and phosphoproteomics of plant mitochondria from non-photosynthetic cells.Front Plant Sci. 2013 Mar 13;4:51. doi: 10.3389/fpls.2013.00051. eCollection 2013. Front Plant Sci. 2013. PMID: 23494127 Free PMC article.
-
Plant mitochondrial pyruvate dehydrogenase complex: purification and identification of catalytic components in potato.Biochem J. 1998 Sep 15;334 ( Pt 3)(Pt 3):571-6. doi: 10.1042/bj3340571. Biochem J. 1998. PMID: 9729464 Free PMC article.
-
Megadalton complexes in the chloroplast stroma of Arabidopsis thaliana characterized by size exclusion chromatography, mass spectrometry, and hierarchical clustering.Mol Cell Proteomics. 2010 Jul;9(7):1594-615. doi: 10.1074/mcp.M000038-MCP201. Epub 2010 Apr 26. Mol Cell Proteomics. 2010. PMID: 20423899 Free PMC article.
-
Primary metabolic pathways and signal transduction in sunflower (Helianthus annuus L.): comparison of transcriptional profiling in leaves and immature embryos using cDNA microarrays.Planta. 2006 Apr;223(5):948-64. doi: 10.1007/s00425-005-0151-6. Epub 2005 Nov 24. Planta. 2006. PMID: 16307285
References
-
- Ali MS, Roche TE, Patel MS. Identification of the essential cysteine residue in the active site of bovine pyruvate dehydrogenase. J Biol Chem. 1993;268:22353–22356. - PubMed
-
- Allen AG, Perham RN, Allison N, Miles JS, Guest JR. Reductive acetylation of tandemly repeated lipoyl domains in the pyruvate dehydrogenase multienzyme complex of Escherichia coli is random order. J Mol Biol. 1989;208:623–633. - PubMed
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Bourguignon J, Macherel D, Neuburger M, Douce R. Isolation, characterization, and sequence analysis of a cDNA clone encoding L-protein, the dihydrolipoamide dehydrogenase component of the glycine cleavage system from pea-leaf mitochondria. Eur J Biochem. 1992;204:865–873. - PubMed
LinkOut - more resources
Full Text Sources

