The herpes simplex virus type 1 U(L)17 gene encodes virion tegument proteins that are required for cleavage and packaging of viral DNA
- PMID: 9557660
- PMCID: PMC109600
- DOI: 10.1128/JVI.72.5.3779-3788.1998
The herpes simplex virus type 1 U(L)17 gene encodes virion tegument proteins that are required for cleavage and packaging of viral DNA
Abstract
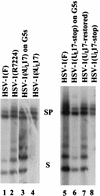
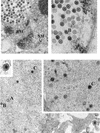
Previous studies have suggested that the U(L)17 gene of herpes simplex virus type 1 (HSV-1) is essential for virus replication. In this study, viral mutants incorporating either a lacZ expression cassette in place of 1,490 bp of the 2,109-bp U(L)17 open reading frame [HSV-1(deltaU(L)17)] or a DNA oligomer containing an in-frame stop codon inserted 778 bp from the 5' end of the U(L)17 open reading frame [HSV-1(U(L)17-stop)] were plaque purified on engineered cell lines containing the U(L)17 gene. A virus derived from HSV-1(U(L)17-stop) but containing a restored U(L)17 gene was also constructed and was designated HSV-1(U(L)17-restored). The latter virus formed plaques and cleaved genomic viral DNA in a manner indistinguishable from wild-type virus. Neither HSV-1(deltaU(L)17) nor HSV-1(U(L)17-stop) formed plaques or produced infectious progeny when propagated on noncomplementing Vero cells. Furthermore, genomic end-specific restriction fragments were not detected in DNA purified from noncomplementing cells infected with HSV-1(deltaU(L)17) or HSV-1(U(L)17-stop), whereas end-specific fragments were readily detected when the viruses were propagated on complementing cells. Electron micrographs of thin sections of cells infected with HSV-1(deltaU(L)17) or HSV-1(U(L)17-stop) illustrated that empty capsids accumulated in the nuclei of Vero cells, whereas DNA-containing capsids accumulated in the nuclei of complementing cells and enveloped virions were found in the cytoplasm and extracellular space. Additionally, protein profiles of capsids purified from cells infected with HSV-1(deltaU(L)17) compared to wild-type virus show no detectable differences. These data indicate that the U(L)17 gene is essential for virus replication and is required for cleavage and packaging of viral DNA. To characterize the U(L)17 gene product, an anti-U(L)17 rabbit polyclonal antiserum was produced. The antiserum reacted strongly with a major protein of apparent Mr 77,000 and weakly with a protein of apparent Mr 72,000 in wild-type infected cell lysates and in virions. Bands of similar sizes were also detected in electrophoretically separated tegument fractions of virions and light particles and yielded tryptic peptides of masses characteristic of the predicted U(L)17 protein. We therefore conclude that the U(L)17 gene products are associated with the virion tegument and note that they are the first tegument-associated proteins shown to be required for cleavage and packaging of viral DNA.
Figures






Similar articles
-
The U(L)15 gene of herpes simplex virus type 1 contains within its second exon a novel open reading frame that is translated in frame with the U(L)15 gene product.J Virol. 1997 Apr;71(4):2666-73. doi: 10.1128/JVI.71.4.2666-2673.1997. J Virol. 1997. PMID: 9060619 Free PMC article.
-
Herpes simplex virus DNA cleavage and packaging: association of multiple forms of U(L)15-encoded proteins with B capsids requires at least the U(L)6, U(L)17, and U(L)28 genes.J Virol. 1998 Apr;72(4):3045-50. doi: 10.1128/JVI.72.4.3045-3050.1998. J Virol. 1998. PMID: 9525627 Free PMC article.
-
A null mutation in the gene encoding the herpes simplex virus type 1 UL37 polypeptide abrogates virus maturation.J Virol. 2001 Nov;75(21):10259-71. doi: 10.1128/JVI.75.21.10259-10271.2001. J Virol. 2001. PMID: 11581394 Free PMC article.
-
Involvement of Terminase Complex in Herpes Simplex Virus Mature Virion Egress.Curr Protein Pept Sci. 2022;23(2):105-113. doi: 10.2174/1389203723666220217144432. Curr Protein Pept Sci. 2022. PMID: 35176987 Review.
-
Comprehensive Analysis of the Tegument Proteins Involved in Capsid Transport and Virion Morphogenesis of Alpha, Beta and Gamma Herpesviruses.Viruses. 2023 Oct 6;15(10):2058. doi: 10.3390/v15102058. Viruses. 2023. PMID: 37896835 Free PMC article. Review.
Cited by
-
Novel class of thiourea compounds that inhibit herpes simplex virus type 1 DNA cleavage and encapsidation: resistance maps to the UL6 gene.J Virol. 2000 Oct;74(19):9054-61. doi: 10.1128/jvi.74.19.9054-9061.2000. J Virol. 2000. PMID: 10982350 Free PMC article.
-
The capsid protein encoded by U(L)17 of herpes simplex virus 1 interacts with tegument protein VP13/14.J Virol. 2010 Aug;84(15):7642-50. doi: 10.1128/JVI.00277-10. Epub 2010 May 26. J Virol. 2010. PMID: 20504930 Free PMC article.
-
How to improve the chemical disinfection of contaminated surfaces by viruses, bacteria and fungus?Eur J Pharm Sci. 2020 Dec 1;155:105559. doi: 10.1016/j.ejps.2020.105559. Epub 2020 Sep 17. Eur J Pharm Sci. 2020. PMID: 32950617 Free PMC article.
-
Herpes simplex virus 1 DNA packaging proteins encoded by UL6, UL15, UL17, UL28, and UL33 are located on the external surface of the viral capsid.J Virol. 2006 Nov;80(21):10894-9. doi: 10.1128/JVI.01364-06. Epub 2006 Aug 18. J Virol. 2006. PMID: 16920825 Free PMC article.
-
Herpes simplex virus type 1 cleavage and packaging proteins UL15 and UL28 are associated with B but not C capsids during packaging.J Virol. 1998 Sep;72(9):7428-39. doi: 10.1128/JVI.72.9.7428-7439.1998. J Virol. 1998. PMID: 9696839 Free PMC article.
References
-
- Addison C, Rixon F J, Palfreyman J W, O’Hara M, Preston V G. Characterisation of a herpes simplex virus type 1 mutant which has a temperature-sensitive defect in penetration of cells and assembly of capsids. Virology. 1984;138:246–259. - PubMed
-
- Addison C, Rixon F J, Preston V G. Herpes simplex virus type 1 UL28 gene product is important for the formation of mature capsids. J Gen Virol. 1990;71:2377–2384. - PubMed
-
- Al-Kobashi M F, Rixon F J, McDougall I, Preston V G. The herpes simplex virus UL33 gene product is required for the assembly of full capsids. Virology. 1991;180:380–388. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

