A promoter region mutation affecting replication of the Tetrahymena ribosomal DNA minichromosome
- PMID: 9566921
- PMCID: PMC110681
- DOI: 10.1128/MCB.18.5.3021
A promoter region mutation affecting replication of the Tetrahymena ribosomal DNA minichromosome
Abstract
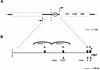
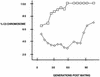
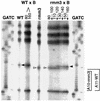
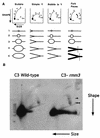
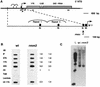
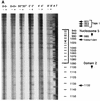
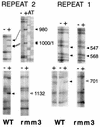
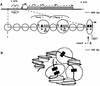
In the ciliated protozoan Tetrahymena thermophila the ribosomal DNA (rDNA) minichromosome replicates partially under cell cycle control and is also subject to a copy number control mechanism. The relationship between rDNA replication and rRNA gene transcription was investigated by the analysis of replication, transcription, and DNA-protein interactions in a mutant rDNA, the rmm3 rDNA. The rmm3 (for rDNA maturation or maintenance mutant 3) rDNA contains a single-base deletion in the rRNA promoter region, in a phylogenetically conserved sequence element that is repeated in the replication origin region of the rDNA minichromosome. The multicopy rmm3 rDNA minichromosome has a maintenance defect in the presence of a competing rDNA allele in heterozygous cells. No difference in the level of rRNA transcription was found between wild-type and rmm3 strains. However, rmm3 rDNA replicating intermediates exhibited an enhanced pause in the region of the replication origin, roughly 750 bp upstream from the rmm3 mutation. In footprinting of isolated nuclei, the rmm3 rDNA lacked the wild-type dimethyl sulfate (DMS) footprint in the promoter region adjacent to the base change. In addition, a DMS footprint in the origin region was lost in the rmm3 rDNA minichromosome. This is the first reported correlation in this system between an rDNA minichromosome maintenance defect and an altered footprint in the origin region. Our results suggest that a promoter region mutation can affect replication without detectably affecting transcription. We propose a model in which interactions between promoter and origin region complexes facilitate replication and maintenance of the Tetrahymena rDNA minichromosome.
Figures

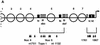
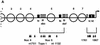
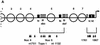

Similar articles
-
Regulatory sequences for the amplification and replication of the ribosomal DNA minichromosome in Tetrahymena thermophila.Mol Cell Biol. 1997 Dec;17(12):7237-47. doi: 10.1128/MCB.17.12.7237. Mol Cell Biol. 1997. PMID: 9372956 Free PMC article.
-
A single-stranded DNA binding protein that specifically recognizes cis-acting sequences in the replication origin and transcriptional promoter region of Tetrahymena rDNA.Biochemistry. 1995 Apr 11;34(14):4583-92. doi: 10.1021/bi00014a011. Biochemistry. 1995. PMID: 7718560
-
Type I elements mediate replication fork pausing at conserved upstream sites in the Tetrahymena thermophila ribosomal DNA minichromosome.Mol Cell Biol. 1997 Aug;17(8):4517-25. doi: 10.1128/MCB.17.8.4517. Mol Cell Biol. 1997. PMID: 9234709 Free PMC article.
-
Developmentally regulated processing and replication of the Tetrahymena rDNA minichromosome.Curr Opin Genet Dev. 1993 Oct;3(5):730-5. doi: 10.1016/s0959-437x(05)80091-7. Curr Opin Genet Dev. 1993. PMID: 8274855 Review.
-
The ribosomal RNA genes of Tetrahymena: structure and function.Eur J Cell Biol. 1985 Jan;36(1):133-51. Eur J Cell Biol. 1985. PMID: 3884336 Review.
Cited by
-
alpha-tubulin minichromosome promoters in the stichotrichous ciliate Stylonychia lemnae.Eukaryot Cell. 2007 Jan;6(1):28-36. doi: 10.1128/EC.00003-06. Epub 2006 Nov 3. Eukaryot Cell. 2007. PMID: 17085637 Free PMC article.
-
A paradoxical mutant GATA factor.Eukaryot Cell. 2004 Apr;3(2):393-405. doi: 10.1128/EC.3.2.393-405.2004. Eukaryot Cell. 2004. PMID: 15075269 Free PMC article.
-
Differential targeting of Tetrahymena ORC to ribosomal DNA and non-rDNA replication origins.EMBO J. 2009 Feb 4;28(3):223-33. doi: 10.1038/emboj.2008.282. Epub 2009 Jan 15. EMBO J. 2009. PMID: 19153611 Free PMC article.
-
Differential use of multiple replication origins in the ribosomal DNA episome of the protozoan parasite Entamoeba histolytica.Nucleic Acids Res. 2003 Apr 15;31(8):2035-44. doi: 10.1093/nar/gkg320. Nucleic Acids Res. 2003. PMID: 12682354 Free PMC article.
-
Three different proteins recognize a multifunctional determinant that controls replication initiation, fork arrest and transcription in Tetrahymena.Nucleic Acids Res. 2000 Feb 1;28(3):843-51. doi: 10.1093/nar/28.3.843. Nucleic Acids Res. 2000. PMID: 10637338 Free PMC article.
References
-
- Blackburn, E., D. Larson, and E. Orias. Unpublished data.
-
- Bonven B J, Gocke E, Westergaard O. A high affinity topoisomerase I binding sequence is clustered at DNAase I hypersensitive sites in Tetrahymena R-chromatin. Cell. 1985;41:541–551. - PubMed
-
- Brewer B J, Fangman W L. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell. 1987;51:463–471. - PubMed
-
- Brown T. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons; 1993. pp. 2.9.1–2.9.6.
-
- Budarf M L, Blackburn E H. Chromatin structure of the telomeric region and 3′-nontranscribed spacer of Tetrahymena ribosomal RNA genes. J Biol Chem. 1986;261:363–369. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
