Cloning and sequencing of a Candida albicans catalase gene and effects of disruption of this gene
- PMID: 9573075
- PMCID: PMC108149
- DOI: 10.1128/IAI.66.5.1953-1961.1998
Cloning and sequencing of a Candida albicans catalase gene and effects of disruption of this gene
Abstract
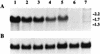
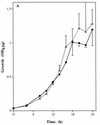
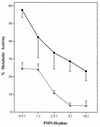
Catalase plays a key role as an antioxidant, protecting aerobic organisms from the toxic effects of hydrogen peroxide, and in some cases has been postulated to be a virulence factor. To help elucidate the function of catalase in Candida albicans, a single C. albicans-derived catalase gene, designated CAT1, was isolated and cloned. Degenerate PCR primers based on highly conserved areas of other fungal catalase genes were used to amplify a 411-bp product from genomic DNA of C. albicans ATCC 10261. By using this product as a probe, catalase clones were isolated from genomic libraries of C. albicans. Nucleotide sequence analysis revealed an open reading frame encoding a protein of 487 amino acid residues. Construction of a CAT1-deficient mutant was achieved by using the Ura-blaster technique for sequential disruption of multiple alleles by integrative transformation using URA3 as a selectable marker. Resulting mutants exhibited normal morphology and comparable growth rates of both yeast and mycelial forms. Enzymatic analysis revealed an abundance of catalase in the wild-type strain but decreasing catalase activity in heterozygous mutants and no detectable catalase in a homozygous null mutant. In vitro assays showed the mutant strains to be more sensitive to damage by both neutrophils and concentrations of exogenous peroxide that were sublethal for the parental strain. Compared to the parental strain, the homozygous null mutant strain was far less virulent for mice in an intravenous infection model of disseminated candidiasis. Definitive linkage of CAT1 with virulence would require restoration of activity by reintroduction of the gene into mutants. However, initial results in mice, taken together with the enhanced susceptibility of catalase-deficient hyphae to damage by human neutrophils, suggest that catalase may enhance the pathogenicity of C. albicans.
Figures







Similar articles
-
Disruption of the human pathogenic yeast Candida albicans catalase gene decreases survival in mouse-model infection and elevates susceptibility to higher temperature and to detergents.Microbiol Immunol. 2003;47(6):395-403. doi: 10.1111/j.1348-0421.2003.tb03376.x. Microbiol Immunol. 2003. PMID: 12906099
-
Cloning and disruption of caPLB1, a phospholipase B gene involved in the pathogenicity of Candida albicans.J Biol Chem. 1998 Oct 2;273(40):26078-86. doi: 10.1074/jbc.273.40.26078. J Biol Chem. 1998. PMID: 9748287
-
A hyphal-specific chitin synthase gene (CHS2) is not essential for growth, dimorphism, or virulence of Candida albicans.Proc Natl Acad Sci U S A. 1994 Jun 21;91(13):6216-20. doi: 10.1073/pnas.91.13.6216. Proc Natl Acad Sci U S A. 1994. PMID: 8016141 Free PMC article.
-
URA3 as a selectable marker for disruption and virulence assessment of Candida albicans genes.Trends Microbiol. 2003 Feb;11(2):69-73. doi: 10.1016/s0966-842x(02)00029-x. Trends Microbiol. 2003. PMID: 12598128 Review.
-
Extracellular phospholipases as universal virulence factor in pathogenic fungi.Nihon Ishinkin Gakkai Zasshi. 1998;39(2):55-9. doi: 10.3314/jjmm.39.55. Nihon Ishinkin Gakkai Zasshi. 1998. PMID: 9580028 Review.
Cited by
-
Regulation of the transcriptional response to oxidative stress in fungi: similarities and differences.Eukaryot Cell. 2003 Jun;2(3):381-9. doi: 10.1128/EC.2.3.381-389.2003. Eukaryot Cell. 2003. PMID: 12796283 Free PMC article. Review. No abstract available.
-
Enterococcus faecalis heme-dependent catalase.J Bacteriol. 2002 Nov;184(22):6351-6. doi: 10.1128/JB.184.22.6351-6356.2002. J Bacteriol. 2002. PMID: 12399505 Free PMC article.
-
Combining Genome-Wide Gene Expression Analysis (RNA-seq) and a Gene Editing Platform (CRISPR-Cas9) to Uncover the Selectively Pro-oxidant Activity of Aurone Compounds Against Candida albicans.Front Microbiol. 2021 Jul 15;12:708267. doi: 10.3389/fmicb.2021.708267. eCollection 2021. Front Microbiol. 2021. PMID: 34335543 Free PMC article.
-
Zinc Cluster Transcription Factors Alter Virulence in Candida albicans.Genetics. 2017 Feb;205(2):559-576. doi: 10.1534/genetics.116.195024. Epub 2016 Dec 7. Genetics. 2017. PMID: 27932543 Free PMC article.
-
Role of the Hog1 stress-activated protein kinase in the global transcriptional response to stress in the fungal pathogen Candida albicans.Mol Biol Cell. 2006 Feb;17(2):1018-32. doi: 10.1091/mbc.e05-06-0501. Epub 2005 Dec 7. Mol Biol Cell. 2006. PMID: 16339080 Free PMC article.
References
-
- Beck-Sague C M, Jarvis W R. National nosocomial infections surveillance system. Secular trends in the epidemiology of nosocomial fungal infections in the United States, 1980–1990. J Infect Dis. 1993;167:1247–1251. - PubMed
-
- Cohen G, Rapatz W, Ruiz H. Sequence of the Saccharomyces cerevisiae CTA1 gene and amino acid sequence of catalase A derived from it. Eur J Biochem. 1988;176:159–163. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

