Mutations in the leucine zipper-like heptad repeat sequence of human immunodeficiency virus type 1 gp41 dominantly interfere with wild-type virus infectivity
- PMID: 9573241
- PMCID: PMC110011
- DOI: 10.1128/JVI.72.6.4765-4774.1998
Mutations in the leucine zipper-like heptad repeat sequence of human immunodeficiency virus type 1 gp41 dominantly interfere with wild-type virus infectivity
Abstract
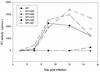
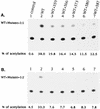
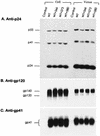
It has been previously shown that a proline substitution for any of the conserved leucine or isoleucine residues located in the leucine zipper-like heptad repeat sequence of human immunodeficiency virus type 1 (HIV-1) gp41 renders viruses noninfectious and envelope (Env) protein unable to mediate membrane fusion (S. S.-L. Chen, C.-N. Lee, W.-R. Lee, K. McIntosh, and T.-M. Lee, J. Virol. 67:3615-3619, 1993; S. S.-L. Chen, J. Virol. 68:2002-2010, 1994). To understand whether these variants could act as trans-dominant inhibitory mutants, the ability of these mutants to inhibit wild-type (wt) virus infectivity was examined. Comparable amounts of cell- and virion-associated gag gene products as well as virion-associated gp41 were found in transfection with wt or mutant HIV-1 provirus. Viruses obtained from coexpression of wt provirus with mutant 566 or 580 provirus inhibited more potently the production of infectious virus than did viruses generated from cotransfection of wt provirus with other mutant proviruses. Nevertheless, all viruses produced from mixed transfection showed decreased infectivity compared with that of the wt virus when a multinuclear-activation beta-galactosidase induction assay was performed. The ability of wt Env to induce cytopathic effects was inhibited by coexpression with mutant Env. Coexpression of mutants inhibited the ability of the wt protein to mediate virus-to-cell transmission, as demonstrated by an env trans-complementation assay with a defective HIV-1 proviral vector. These observations indicated that mutant Env, per se, interferes with wt Env function. Moreover, cotransfection of wt and mutant proviruses produced amounts of cell- and virion-associated gag gene products comparable to those produced by transfection of wt provirus. Similar amounts of gp41 were also found in virions generated from wt-mutant cotransfection as well as from wt transfection alone. These results indicated that the inhibitory effect conferred by mutants on the wt virus infectivity does not involve the late steps of Gag protein assembly and budding, but they suggest that the wt and mutant Env proteins form a dysfunctional hetero-oligomer which is impaired in an early step of the virus replication cycle. Our study demonstrates that mutations in the HIV-1 gp41 leucine zipper-like heptad repeat sequence dominantly inhibit infectious virus production.
Figures






References
-
- Abacioglu Y H, Fouts T R, Laman J D, Claasen E, Pincus S H, Moore J P, Roby C A, Kamin-Lewis R, Lewis G K. Epitope mapping and topology of baculovirus-expressed HIV-1 gp160 determined with a panel of murine monoclonal antibodies. AIDS Res Hum Retroviruses. 1994;10:371–381. - PubMed
-
- Bates P. Chemokine receptors and HIV-1: an attractive pair? Cell. 1996;86:1–3. - PubMed
-
- Blacklow S C, Lu M, Kim P S. A trimeric subdomain of the simian immunodeficiency virus envelope glycoprotein. Biochemistry. 1995;34:14955–14962. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

