The competence transcription factor of Bacillus subtilis recognizes short A/T-rich sequences arranged in a unique, flexible pattern along the DNA helix
- PMID: 9585513
- PMCID: PMC316842
- DOI: 10.1101/gad.12.10.1539
The competence transcription factor of Bacillus subtilis recognizes short A/T-rich sequences arranged in a unique, flexible pattern along the DNA helix
Abstract
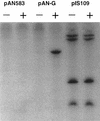
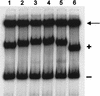
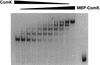
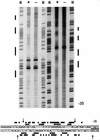
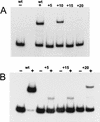
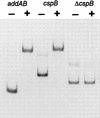
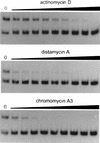
The development of genetic competence in Bacillus subtilis is regulated by a complex signal transduction cascade, which leads to the synthesis of the competence transcription factor (CTF). Previous studies suggested that CTF is encoded by comK. ComK is required for the transcription of comK itself, as well as of the late competence genes encoding the DNA uptake machinery and of genes required for homologous recombination. Here, we used purified ComK to study its role in transcription and to determine the DNA recognition sequence for ComK. In vitro transcription from the comG promoter, which depends on ComK in vivo, was observed on the addition of purified ComK together with Bacillus subtilis RNA polymerase, proving that ComK is CTF. To determine the DNA sequences involved in ComK recognition, footprinting analysis was performed with promoter fragments of the CTF-dependent genes: comC, comE, comF, comG, comK, and addAB. The ComK binding sites determined by DNase I protection experiments were unusually long, with average lengths of approximately 65 bp, and displayed only weak sequence similarities. Hydroxy-radical footprinting, performed with the addAB promoter, revealed a unique arrangement of four short A/T-rich sequences. Gel retardation experiments indicated that four molecules of ComK bound the addAB promoter and the dyad symmetrical arrangement of the four A/T-rich sequences implied that ComK functions as a tetramer composed of two dimers each recognizing the motif AAAAN5TTTT. Comparable A/T-rich sequences were identified in all six DNase I footprints and could be used to predict ComK targets in the B. subtilis genome. On the basis of the variability in distance between the ComK-dimer binding sites, ComK-regulated promoters could be divided into three classes, demonstrating a remarkable flexibility in the binding of ComK. The pattern of hydroxy-radical protections suggested that ComK binds at one face of the DNA helix through the minor groove. This inference was strengthened by the observation that minor groove binding drugs inhibited the binding of ComK.
Figures
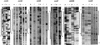


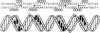

Similar articles
-
comK encodes the competence transcription factor, the key regulatory protein for competence development in Bacillus subtilis.Mol Microbiol. 1995 Feb;15(3):455-62. doi: 10.1111/j.1365-2958.1995.tb02259.x. Mol Microbiol. 1995. PMID: 7783616
-
The Bacillus subtilis transition state regulator AbrB binds to the -35 promoter region of comK.FEMS Microbiol Lett. 2003 Jan 28;218(2):299-304. doi: 10.1111/j.1574-6968.2003.tb11532.x. FEMS Microbiol Lett. 2003. PMID: 12586407
-
Mechanism of transcription activation at the comG promoter by the competence transcription factor ComK of Bacillus subtilis.J Bacteriol. 2004 Feb;186(4):1120-8. doi: 10.1128/JB.186.4.1120-1128.2004. J Bacteriol. 2004. PMID: 14762007 Free PMC article.
-
Genetic competence in Bacillus subtilis.Microbiol Rev. 1991 Sep;55(3):395-424. doi: 10.1128/mr.55.3.395-424.1991. Microbiol Rev. 1991. PMID: 1943994 Free PMC article. Review.
-
Controlling competence in Bacillus subtilis: shared use of regulators.Microbiology (Reading). 2003 Jan;149(Pt 1):9-17. doi: 10.1099/mic.0.26003-0. Microbiology (Reading). 2003. PMID: 12576575 Review.
Cited by
-
The ResD response regulator, through functional interaction with NsrR and fur, plays three distinct roles in Bacillus subtilis transcriptional control.J Bacteriol. 2014 Jan;196(2):493-503. doi: 10.1128/JB.01166-13. Epub 2013 Nov 8. J Bacteriol. 2014. PMID: 24214949 Free PMC article.
-
High-Throughput Genetic Screens Identify a Large and Diverse Collection of New Sporulation Genes in Bacillus subtilis.PLoS Biol. 2016 Jan 6;14(1):e1002341. doi: 10.1371/journal.pbio.1002341. eCollection 2016 Jan. PLoS Biol. 2016. PMID: 26735940 Free PMC article.
-
Bistability in the Bacillus subtilis K-state (competence) system requires a positive feedback loop.Mol Microbiol. 2005 May;56(3):615-24. doi: 10.1111/j.1365-2958.2005.04592.x. Mol Microbiol. 2005. PMID: 15819619 Free PMC article.
-
The MogR transcriptional repressor regulates nonhierarchal expression of flagellar motility genes and virulence in Listeria monocytogenes.PLoS Pathog. 2006 Apr;2(4):e30. doi: 10.1371/journal.ppat.0020030. Epub 2006 Apr 14. PLoS Pathog. 2006. PMID: 16617375 Free PMC article.
-
Regulation of ykrL (htpX) by Rok and YkrK, a novel type of regulator in Bacillus subtilis.J Bacteriol. 2012 Jun;194(11):2837-45. doi: 10.1128/JB.00324-12. Epub 2012 Mar 23. J Bacteriol. 2012. PMID: 22447908 Free PMC article.
References
-
- Dubnau D. Genetic exchange and homologous recombination. In: Sonenshein AL, Hoch JA, Losick R, editors. Bacillus subtilis and other Gram-positive bacteria. Washington, D.C.: American Society for Microbiology; 1993. pp. 555–584.
-
- Duplay P, Bedouelle H, Fowler A, Zabin I, Saurin W, Hofnung M. Sequences of the malE gene and of its product, the maltose-binding protein of Escherichia coli K12. J Biol Chem. 1984;259:10606–10613. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
