A spore coat protein, CotS, of Bacillus subtilis is synthesized under the regulation of sigmaK and GerE during development and is located in the inner coat layer of spores
- PMID: 9603889
- PMCID: PMC107266
- DOI: 10.1128/JB.180.11.2968-2974.1998
A spore coat protein, CotS, of Bacillus subtilis is synthesized under the regulation of sigmaK and GerE during development and is located in the inner coat layer of spores
Abstract
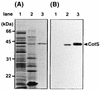
The spore coat of Bacillus subtilis has a unique morphology and consists of polypeptides of different sizes, whose synthesis and assembly are precisely regulated by a cascade of transcription factors and regulatory proteins. We examined the factors that regulate cotS gene expression and CotS assembly into the coat layer of B. subtilis by Northern blot and Western blot analysis. Transcription of cotS mRNA was not detected in sporulating cells of sigmaK and gerE mutants by Northern blot analysis. By Western blot analysis using anti-CotS antibody, CotS was first detected in protein samples solubilized from wild-type cells at 5 h after the start of sporulation. CotS was not detected in the vegetative cells and spores of a gerE mutant or in the spores of mutants deficient in sigmaE, sigmaF, sigmaG, or sigmaK. CotS was detected in the sporangium but not in the spores of a cotE mutant. The sequence of the promoter region of cotS was similar to the consensus sequences for binding of sigmaK and GerE. These results demonstrate that sigmaK and GerE are required for cotS expression and that CotE is essential for the assembly of CotS in the coat. Immunoelectron microscopic observation using anti-CotS antibody revealed that CotS is located within the spore coat, in particular in the inner coats of dormant spores.
Figures







References
-
- Abe A, Ogawa S, Kohno T, Watabe K. Purification of Bacillus subtilis spore coat protein by electrophoretic elution procedure and determination of NH2-terminal amino acid sequences. Microbiol Immunol. 1993;37:809–812. - PubMed
-
- Abe A, Koide H, Kohno T, Watabe K. A Bacillus subtilis spore coat polypeptide gene, cotS. Microbiology. 1995;141:1433–1442. - PubMed
-
- Adam D, Roels S, Beall B, Moran C P, Jr, Losick R. Subcellular localization of proteins involved in the assembly of the spore coat of Bacillus subtilis. Genes Dev. 1994;8:234–244. - PubMed
-
- Aronson A I, Song H-Y, Bourne N. Gene structure and precursor processing of a novel Bacillus subtilis spore coat protein. Mol Microbiol. 1989;3:437–444. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

