Differential selection of cells with proviral c-myc and c-erbB integrations after avian leukosis virus infection
- PMID: 9621008
- PMCID: PMC110195
- DOI: 10.1128/JVI.72.7.5517-5525.1998
Differential selection of cells with proviral c-myc and c-erbB integrations after avian leukosis virus infection
Abstract
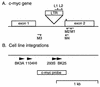
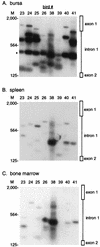
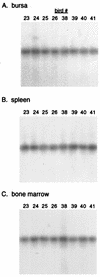
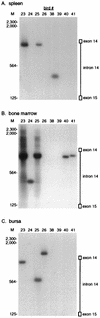
Avian leukosis virus (ALV) infection induces bursal lymphomas in chickens after proviral integration within the c-myc proto-oncogene and induces erythroblastosis after integration within the c-erbB proto-oncogene. A nested PCR assay was used to analyze the appearance of these integrations at an early stage of tumor induction after infection of embryos. Five to eight distinct proviral c-myc integration events were amplified from bursas of infected 35-day-old birds, in good agreement with the number of transformed bursal follicles arising with these integrations. Cells containing these integrations are remarkably common, with an estimated 1 in 350 bursal cells having proviral c-myc integrations. These integrations were clustered within the 3' half of c-myc intron 1, in a pattern similar to that observed in bursal lymphomas. Bone marrow and spleen showed a similar number and pattern of integrations clustered within 3' c-myc intron 1, indicating that this region is a common integration target whether or not that tissue undergoes tumor induction. While all tissues showed equivalent levels of viral infection, cells with c-myc integrations were much more abundant in the bursa than in other tissues, indicating that cells with proviral c-myc integrations are preferentially expanded within the bursal environment. Proviral integration within the c-erbB gene was also analyzed, to detect clustered c-erbB intron 14 integrations associated with erythroblastosis. Proviral c-erbB integrations were equally abundant in the bone marrow, spleen, and bursa. These integrations were randomly situated upstream of c-erbB exon 15, indicating that cells carrying 3' intron 14 integrations must be selected during induction of erythroblastosis.
Figures




Similar articles
-
Resistance to avian leukosis virus lymphomagenesis occurs subsequent to proviral c-myc integration.Oncogene. 1999 Jan 7;18(1):201-9. doi: 10.1038/sj.onc.1202283. Oncogene. 1999. PMID: 9926935
-
Reduced Myc overexpression and normal B-cell differentiation mediate resistance to avian leukosis virus lymphomagenesis.Oncogene. 2004 May 27;23(25):4413-21. doi: 10.1038/sj.onc.1207577. Oncogene. 2004. PMID: 15064748
-
c-erbB activation in avian leukosis virus-induced erythroblastosis: clustered integration sites and the arrangement of provirus in the c-erbB alleles.Proc Natl Acad Sci U S A. 1985 Apr;82(8):2287-91. doi: 10.1073/pnas.82.8.2287. Proc Natl Acad Sci U S A. 1985. PMID: 2986110 Free PMC article.
-
The viral envelope is a major determinant for the induction of lymphoid and myeloid tumours by avian leukosis virus subgroups A and J, respectively.J Gen Virol. 2002 Oct;83(Pt 10):2553-2561. doi: 10.1099/0022-1317-83-10-2553. J Gen Virol. 2002. PMID: 12237439
-
An aberrant avian leukosis virus provirus inserted downstream from the chicken c-myc coding sequence in a bursal lymphoma results from intrachromosomal recombination between two proviruses and deletion of cellular DNA.J Virol. 1987 Jun;61(6):1828-33. doi: 10.1128/JVI.61.6.1828-1833.1987. J Virol. 1987. PMID: 3033315 Free PMC article.
Cited by
-
Acutely transforming avian leukosis virus subgroup J strain 966: defective genome encodes a 72-kilodalton Gag-Myc fusion protein.J Virol. 2001 May;75(9):4219-25. doi: 10.1128/JVI.75.9.4219-4225.2001. J Virol. 2001. PMID: 11287571 Free PMC article.
References
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons, Inc.; 1994.
-
- Bloom S E, Delany M E, Muscarella D E. Constant and variable features of avian chromosomes. In: Etches R J, Gibbons A M V, editors. Manipulation of the avian genome. Boca Raton, Fla: CRC Press, Inc.; 1993. pp. 39–60.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

