A small yeast RNA blocks hepatitis C virus internal ribosome entry site (HCV IRES)-mediated translation and inhibits replication of a chimeric poliovirus under translational control of the HCV IRES element
- PMID: 9621022
- PMCID: PMC110227
- DOI: 10.1128/JVI.72.7.5638-5647.1998
A small yeast RNA blocks hepatitis C virus internal ribosome entry site (HCV IRES)-mediated translation and inhibits replication of a chimeric poliovirus under translational control of the HCV IRES element
Erratum in
- J Virol 1998 Nov;72(11):9419
Abstract
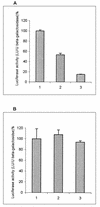
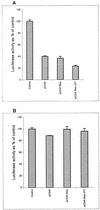
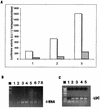
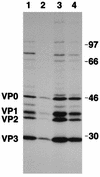
Hepatitis C virus (HCV) infection frequently leads to chronic hepatitis and cirrhosis of the liver and has been linked to development of hepatocellular carcinoma. We previously identified a small yeast RNA (IRNA) capable of specifically inhibiting poliovirus (PV) internal ribosome entry site (IRES)-mediated translation. Here we report that IRNA specifically inhibits HCV IRES-mediated translation both in vivo and in vitro. A number of human hepatoma (Huh-7) cell lines expressing IRNA were prepared and characterized. Constitutive expression of IRNA was not detrimental to cell growth. HCV IRES-mediated cap-independent translation was markedly inhibited in cells constitutively expressing IRNA compared to control hepatoma cells. However, cap-dependent translation was not significantly affected in these cell lines. Additionally, Huh-7 cells constitutively expressing IRNA became refractory to infection by a PV-HCV chimera in which the PV IRES is replaced by the HCV IRES. In contrast, replication of a PV-encephalomyocarditis virus (EMCV) chimera containing the EMCV IRES element was not affected significantly in the IRNA-producing cell line. Finally, the binding of the La autoantigen to the HCV IRES element was specifically and efficiently competed by IRNA. These results provide a basis for development of novel drugs effective against HCV infection.
Figures




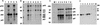
Similar articles
-
A peptide from autoantigen La blocks poliovirus and hepatitis C virus cap-independent translation and reveals a single tyrosine critical for La RNA binding and translation stimulation.J Virol. 2004 Apr;78(7):3763-76. doi: 10.1128/jvi.78.7.3763-3776.2004. J Virol. 2004. PMID: 15016896 Free PMC article.
-
Inhibition of internal entry site (IRES)-mediated translation by a small yeast RNA: a novel strategy to block hepatitis C virus protein synthesis.Front Biosci. 1998 Dec 1;3:D1241-52. doi: 10.2741/a359. Front Biosci. 1998. PMID: 9835647 Review.
-
A small yeast RNA inhibits HCV IRES mediated translation and inhibits replication of poliovirus in vivo.World J Gastroenterol. 2003 May;9(5):1008-13. doi: 10.3748/wjg.v9.i5.1008. World J Gastroenterol. 2003. PMID: 12717847 Free PMC article.
-
Inhibitor RNA blocks the protein translation mediated by hepatitis C virus internal ribosome entry site in vivo.World J Gastroenterol. 2004 Mar 1;10(5):664-7. doi: 10.3748/wjg.v10.i5.664. World J Gastroenterol. 2004. PMID: 14991934 Free PMC article.
-
Targeting internal ribosome entry site (IRES)-mediated translation to block hepatitis C and other RNA viruses.FEMS Microbiol Lett. 2004 May 15;234(2):189-99. doi: 10.1016/j.femsle.2004.03.045. FEMS Microbiol Lett. 2004. PMID: 15135522 Review.
Cited by
-
Role of the 5'-proximal stem-loop structure of the 5' untranslated region in replication and translation of hepatitis C virus RNA.J Virol. 2003 Mar;77(5):3312-8. doi: 10.1128/jvi.77.5.3312-3318.2003. J Virol. 2003. PMID: 12584356 Free PMC article.
-
La autoantigen is necessary for optimal function of the poliovirus and hepatitis C virus internal ribosome entry site in vivo and in vitro.Mol Cell Biol. 2004 Aug;24(15):6861-70. doi: 10.1128/MCB.24.15.6861-6870.2004. Mol Cell Biol. 2004. PMID: 15254251 Free PMC article.
-
A peptide from autoantigen La blocks poliovirus and hepatitis C virus cap-independent translation and reveals a single tyrosine critical for La RNA binding and translation stimulation.J Virol. 2004 Apr;78(7):3763-76. doi: 10.1128/jvi.78.7.3763-3776.2004. J Virol. 2004. PMID: 15016896 Free PMC article.
-
Internal ribosome entry sites (IRESs): reality and use.Transgenic Res. 1999 Jun;8(3):157-77. doi: 10.1023/a:1008909908180. Transgenic Res. 1999. PMID: 10478488 Review.
-
La autoantigen is required for the internal ribosome entry site-mediated translation of Coxsackievirus B3 RNA.Nucleic Acids Res. 2002 Oct 15;30(20):4500-8. doi: 10.1093/nar/gkf583. Nucleic Acids Res. 2002. PMID: 12384597 Free PMC article.
References
-
- Blyn L B, Swiderek K M, Richards O, Stahl D C, Semler B L, Ehrenfeld E. Poly r(C) binding protein 2, binds to stem-loop IV of the poliovirus RNA 5′noncoding region: identification by automated ligand chromatography-tandem mass spectrometry. Proc Natl Acad Sci USA. 1996;93:11115–11120. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

