Structural and antigenic analysis of a truncated form of the herpes simplex virus glycoprotein gH-gL complex
- PMID: 9621073
- PMCID: PMC110415
- DOI: 10.1128/JVI.72.7.6092-6103.1998
Structural and antigenic analysis of a truncated form of the herpes simplex virus glycoprotein gH-gL complex
Abstract
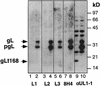
The herpes simplex virus (HSV) gH-gL complex is essential for virus infectivity and is a major antigen for the host immune system. The association of gH with gL is required for correct folding, cell surface trafficking, and membrane presentation of the complex. Previously, a mammalian cell line was constructed which produces a secreted form of gHt-gL complex lacking the transmembrane and cytoplasmic tail regions of gH. gHt-gL retains a conformation similar to that of its full-length counterpart in HSV-infected cells. Here, we examined the structural and antigenic properties of gHt-gL. We first determined its stoichiometry and carbohydrate composition. We found that the complex consists of one molecule each of gH and gL. The N-linked carbohydrate (N-CHO) site on gL and most of the N-CHO sites on gH are utilized, and both proteins also contain O-linked carbohydrate and sialic acid. These results suggest that the complex is processed to the mature form via the Golgi network prior to secretion. To determine the antigenically active sites of gH and gL, we mapped the epitopes of a panel of gH and gL monoclonal antibodies (MAbs), using a series of gH and gL C-terminal truncation variant proteins produced in transiently transfected mammalian cells. Sixteen gH MAbs (including H6 and 37S) reacted with the N-terminal portion of gH between amino acids 19 and 276. One of the gH MAbs, H12, reacted with the middle portion of gH (residues 476 to 678). Nine gL MAbs (including 8H4 and VIII 62) reacted with continuous epitopes within the C-terminal portion of gL, and this region was further mapped within amino acids 168 to 178 with overlapping synthetic peptides. Finally, plasmids expressing the gH and gL truncations were employed in cotransfection assays to define the minimal regions of both gH and gL required for complex formation and secretion. The first 323 amino acids of gH and the first 161 amino acids of gL can form a stable secreted hetero-oligomer with gL and gH792, respectively, while gH323-gL168 is the smallest secreted hetero-oligomer. The first 648 amino acids of gH are required for reactivity with MAbs LP11 and 53S, indicating that a complex of gH648-gL oligomerizes into the correct conformation. The data suggest that both antigenic activity and oligomeric structure require the amino-terminal portions of gH and gL.
Figures



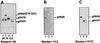

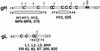

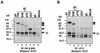

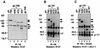

References
-
- Anderson R A, Liu D X, Gompels U A. Definition of a human herpesvirus-6 betaherpesvirus-specific domain in glycoprotein gH that governs interaction with glycoprotein gL: substitution of human cytomegalovirus glycoproteins permits group-specific complex formation. Virology. 1996;217:517–526. - PubMed
-
- Buckmaster E A, Gompels U, Minson A. Characterisation and physical mapping of an HSV-1 glycoprotein of approximately 115 × 103 molecular weight. Virology. 1984;139:408–413. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

