Schistosome satellite DNA encodes active hammerhead ribozymes
- PMID: 9632772
- PMCID: PMC108972
- DOI: 10.1128/MCB.18.7.3880
Schistosome satellite DNA encodes active hammerhead ribozymes
Abstract
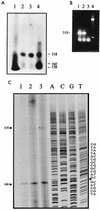
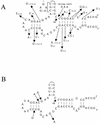
Using a computer program designed to search for RNA structural motifs in sequence databases, we have found a hammerhead ribozyme domain encoded in the Smalpha repetitive DNA of Schistosoma mansoni. Transcripts of these repeats are expressed as long multimeric precursor RNAs that cleave in vitro and in vivo into unit-length fragments. This RNA domain is able to engage in both cis and trans cleavage typical of the hammerhead ribozyme. Further computer analysis of S. mansoni DNA identified a potential trans cleavage site in the gene coding for a synaptobrevin-like protein, and RNA transcribed from this gene was efficiently cleaved by the Smalpha ribozyme in vitro. Similar families of repeats containing the hammerhead domain were found in the closely related Schistosoma haematobium and Schistosomatium douthitti species but were not present in Schistosoma japonicum or Heterobilharzia americana, suggesting that the hammerhead domain was not acquired from a common schistosome ancestor.
Figures





References
-
- Barker S C, Blair D. Molecular phylogeny of Schistosoma species supports traditional groupings within the genus. J Parasitol. 1996;82:292–298. - PubMed
-
- Boeke J D. LINEs and Alus—the poly A connection. Nat Genet. 1997;16:6–7. - PubMed
-
- Bratty J, Chartrand P, Ferbeyre G, Cedergren R. The hammerhead RNA domain, a model ribozyme. Biochim Biophys Acta. 1993;1216:345–359. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
