Adjustment of conformational flexibility is a key event in the thermal adaptation of proteins
- PMID: 9636162
- PMCID: PMC22632
- DOI: 10.1073/pnas.95.13.7406
Adjustment of conformational flexibility is a key event in the thermal adaptation of proteins
Abstract
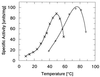
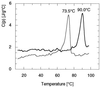
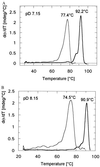
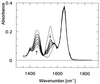
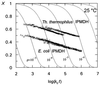
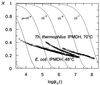
3-Isopropylmalate dehydrogenase (IPMDH, E.C. 1.1.1.85) from the thermophilic bacterium Thermus thermophilus HB8 is homologous to IPMDH from the mesophilic Escherichia coli, but has an approximately 17 degreesC higher melting temperature. Its temperature optimum is 22-25 degreesC higher than that of the E. coli enzyme; however, it is hardly active at room temperature. The increased conformational rigidity required to stabilize the thermophilic enzyme against heat denaturation might explain its different temperature-activity profile. Hydrogen/deuterium exchange studies were performed on this thermophilic-mesophilic enzyme pair to compare their conformational flexibilities. It was found that Th. thermophilus IPMDH is significantly more rigid at room temperature than E. coli IPMDH, whereas the enzymes have nearly identical flexibilities under their respective optimal working conditions, suggesting that evolutionary adaptation tends to maintain a "corresponding state" regarding conformational flexibility. These observations confirm that conformational fluctuations necessary for catalytic function are restricted at room temperature in the thermophilic enzyme, suggesting a close relationship between conformational flexibility and enzyme function.
Figures
Similar articles
-
A better enzyme to cope with cold. Comparative flexibility studies on psychrotrophic, mesophilic, and thermophilic IPMDHs.J Biol Chem. 2001 Jul 27;276(30):28121-5. doi: 10.1074/jbc.M104432200. Epub 2001 May 21. J Biol Chem. 2001. PMID: 11369782
-
Urea-induced unfolding and conformational stability of 3-isopropylmalate dehydrogenase from the Thermophile thermus thermophilus and its mesophilic counterpart from Escherichia coli.Biochemistry. 1999 Jan 26;38(4):1332-7. doi: 10.1021/bi982380v. Biochemistry. 1999. PMID: 9930995
-
High thermal stability of 3-isopropylmalate dehydrogenase from Thermus thermophilus resulting from low DeltaC(p) of unfolding.Protein Eng. 2001 Dec;14(12):961-6. doi: 10.1093/protein/14.12.961. Protein Eng. 2001. PMID: 11809926
-
[Structure of 3-isopropylmalate dehydrogenase from an extreme thermophile--a new model enzyme molecule for the study of structure-function and structure-stability relationships].Tanpakushitsu Kakusan Koso. 1993 Jul;38(9):1453-65. Tanpakushitsu Kakusan Koso. 1993. PMID: 8356215 Review. Japanese. No abstract available.
-
A Minireview on Temperature Dependent Protein Conformational Sampling.Protein J. 2021 Aug;40(4):545-553. doi: 10.1007/s10930-021-10012-x. Epub 2021 Jun 28. Protein J. 2021. PMID: 34181188 Review.
Cited by
-
Comparison of the structural stability and dynamic properties of recombinant anthrax protective antigen and its 2-fluorohistidine-labeled analogue.J Pharm Sci. 2012 Nov;101(11):4118-28. doi: 10.1002/jps.23294. Epub 2012 Aug 21. J Pharm Sci. 2012. PMID: 22911632 Free PMC article.
-
Denaturing action of urea and guanidine hydrochloride towards two thermophilic esterases.Biochem J. 2002 Nov 1;367(Pt 3):857-63. doi: 10.1042/BJ20020695. Biochem J. 2002. PMID: 12160466 Free PMC article.
-
Substrate inhibition imposes fitness penalty at high protein stability.Proc Natl Acad Sci U S A. 2019 Jun 4;116(23):11265-11274. doi: 10.1073/pnas.1821447116. Epub 2019 May 16. Proc Natl Acad Sci U S A. 2019. PMID: 31097595 Free PMC article.
-
Living on the Edge: Physiological and Kinetic Trade-Offs Shape Thermal Tolerance in Intertidal Crabs From Tropical to Sub-Antarctic South America.Front Physiol. 2020 Apr 24;11:312. doi: 10.3389/fphys.2020.00312. eCollection 2020. Front Physiol. 2020. PMID: 32390860 Free PMC article.
-
Dynamics of thermodynamically stable, kinetically trapped, and inhibitor-bound states of pepsin.Biophys J. 2011 Oct 5;101(7):1699-709. doi: 10.1016/j.bpj.2011.08.002. Biophys J. 2011. PMID: 21961596 Free PMC article.
References
-
- Careri G, Fasella P, Gratton E. CRC Crit Rev Biochem. 1975;3:141–164. - PubMed
-
- Cooper A. Prog Biophys Mol Biol. 1984;34:181–214. - PubMed
-
- Linderström-Lang K M, Schellman J A. In: The Enzymes. Boyer P, editor. New York: Academic; 1959. pp. 443–510.
-
- Straub F B. Adv Enzymol. 1964;2:89–114. - PubMed
-
- Hvidt A, Nielsen S O. Adv Protein Chem. 1966;21:287–386. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

