Binary sibling neuronal cell fate decisions in the Drosophila embryonic central nervous system are nonstochastic and require inscuteable-mediated asymmetry of ganglion mother cells
- PMID: 9637687
- PMCID: PMC316905
- DOI: 10.1101/gad.12.12.1858
Binary sibling neuronal cell fate decisions in the Drosophila embryonic central nervous system are nonstochastic and require inscuteable-mediated asymmetry of ganglion mother cells
Abstract
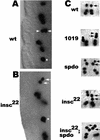
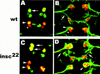
Asymmetric cell division is a widespread mechanism in developing tissues that leads to the generation of cell diversity. In the embryonic central nervous system of Drosophila melanogaster, secondary precursor cells-ganglion mother cells (GMCs)-divide and produce postmitotic neurons that take on different cell fates. In this study, we show that binary fate decision of two pairs of sibling neurons is accomplished through the interplay of Notch (N) signaling and the intrinsic fate determinant Numb. We show that GMCs have apical-basal polarity and Numb localization and the orientation of division are coordinated to segregate Numb to only one sibling cell. The correct positioning of Numb and the proper orientation of division require Inscuteable (Insc). Loss of insc results in the generation of equivalent sibling cells. Our results provide evidence that sibling neuron fate decision is nonstochastic and normally depends on the presence of Numb in one of the two siblings. Moreover, our data suggest that the fate of some sibling neurons may be regulated by signals that do not require lateral interaction between the sibling cells.
Figures






References
-
- Artavanis-Tsakonas A, Matsuno K, Fortini ME. Notch signaling. Science. 1995;268:225–232. - PubMed
-
- Bossing T, Udolph G, Doe CQ, Technau G. The embryonic CNS lineages of Drosophila melanogaster: I. Neuroblast lineages derived from the ventral half of the neuroectoderm. Dev Biol. 1996;179:41–64. - PubMed
-
- Burchard S, Paululat A, Hinz U, Renkawitz-Pohl R. The mutant not enough muscles (nem) reveals reduction of the Drosophila embryonic muscle pattern. J Cell Sci. 1995;108:1443–1454. - PubMed
-
- Campos-Ortega JA. Numb diverts Notch pathway off the Tramtrack. Neuron. 1996;17:1–4. - PubMed
-
- Campos-Ortega JA, Hartenstein V. The embryonic development of Drosophila melanogaster. Berlin, Germany: Springer-Verlag; 1985.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
