Molecular population genetics of the Arabidopsis CAULIFLOWER regulatory gene: nonneutral evolution and naturally occurring variation in floral homeotic function
- PMID: 9653152
- PMCID: PMC20941
- DOI: 10.1073/pnas.95.14.8130
Molecular population genetics of the Arabidopsis CAULIFLOWER regulatory gene: nonneutral evolution and naturally occurring variation in floral homeotic function
Abstract
The evolution of interspecies differences in morphology requires sufficient within-species variation in developmental regulatory systems on which evolutionary forces can act. Molecular analyses of naturally occurring alleles of the Arabidopsis thaliana CAULIFLOWER locus reveal considerable intraspecific diversity at this floral homeotic gene, and the McDonald-Kreitman test suggests that this gene is evolving in a nonneutral fashion, with an excess of intraspecific replacement polymorphisms. The naturally occurring molecular variation within this floral regulatory gene is associated with functionally different alleles, which can be distinguished phenotypically by their differential ability to direct floral meristem development.
Figures



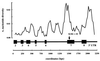
References
-
- Palopoli M F, Patel N H. Curr Opin Gen Dev. 1996;6:502–508. - PubMed
-
- Gould S J. Ontogeny and Phylogeny. Cambridge, MA: Harvard Univ. Press; 1977.
-
- Liljegren S J, Yanofsky M F. Curr Opin Cell Biol. 1996;8:865–869. - PubMed
-
- Weigel D, Meyerowitz E M. Cell. 1994;78:203–209. - PubMed
-
- Coen, E. S. & Nugent, J. M. (1994) Dev. Suppl. 107–116.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

