3'-Processed mRNA is preferentially translated in Chlamydomonas reinhardtii chloroplasts
- PMID: 9671470
- PMCID: PMC109046
- DOI: 10.1128/MCB.18.8.4605
3'-Processed mRNA is preferentially translated in Chlamydomonas reinhardtii chloroplasts
Abstract
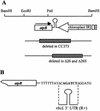
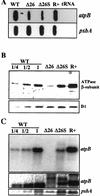
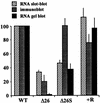
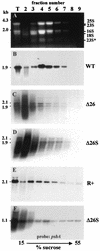
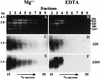
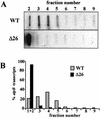
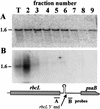
3'-end processing of nucleus-encoded mRNAs includes the addition of a poly(A) tail that is important for translation initiation. Since the vast majority of chloroplast mRNAs acquire their 3' termini by processing yet are not polyadenylated, we asked whether 3' end maturation plays a role in chloroplast translation. A general characteristic of the 3' untranslated regions of chloroplast mRNAs is an inverted repeat (IR) sequence that can fold into a stem-loop structure. These stem-loops and their flanking sequences serve as RNA 3'-end formation signals. Deletion of the Chlamydomonas chloroplast atpB 3' IR in strain Delta26 results in reduced accumulation of atpB transcripts and the chloroplast ATPase beta-subunit, leading to weakly photosynthetic growth. Of the residual atpB mRNA in Delta26, approximately 1% accumulates as a discrete RNA of wild-type size, while the remainder is heterogeneous in length due to the lack of normal 3' end maturation. In this work, we have analyzed whether these unprocessed atpB transcripts are actively translated in vivo. We found that only the minority population of discrete transcripts of wild-type size is associated with polysomes and thus accounts for the ATPase beta-subunit which accumulates in Delta26. Analysis of chloroplast rbcL mRNA revealed that transcripts extending beyond the mature 3' end were not polysome associated. These results suggest that 3'-end processing of chloroplast mRNA is required for or strongly stimulates its translation.
Figures
References
-
- Audren H, Bisanz-Seyer C, Briat J F, Mache R. Structure and transcription of the 5S ribosomal RNA gene from spinach chloroplasts. Curr Genet. 1987;12:263–270. - PubMed
-
- Blowers A D, Klein U, Ellmore G S, Bogorad L. Functional in vivo analyses of the 3′ flanking sequences of the Chlamydomonas chloroplast rbcL and psaB genes. Mol Gen Genet. 1993;238:339–349. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
