Systematic identification of essential genes by in vitro mariner mutagenesis
- PMID: 9671781
- PMCID: PMC21179
- DOI: 10.1073/pnas.95.15.8927
Systematic identification of essential genes by in vitro mariner mutagenesis
Abstract
Although the complete DNA sequences of several microbial genomes are now available, nearly 40% of the putative genes lack identifiable functions. Comprehensive screens and selections for identifying functional classes of genes are needed to convert sequence data into meaningful biological information. One particularly significant group of bacterial genes consists of those that are essential for growth or viability. Here, we describe a simple system for performing transposon mutagenesis on naturally transformable organisms along with a technique to rapidly identify essential or conditionally essential DNA segments. We show the general utility of this approach by applying it to two human pathogens, Haemophilus influenzae and Streptococcus pneumoniae, in which we detected known essential genes and assigned essentiality to several ORFs of unknown function.
Figures
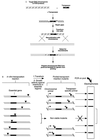
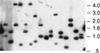


References
-
- Harris S D, Cheng J, Pugh T A, Pringle J R. J Mol Biol. 1992;225:53–65. - PubMed
-
- Gaiano N, Amsterdam A, Kawakami K, Allende M, Becker T, Hopkins N. Nature (London) 1996;383:829–832. - PubMed
-
- Murphy C K, Stewart E J, Beckwith J. Gene. 1995;155:1–7. - PubMed
-
- Blattner F R, Plunkett G R, Bloch C A, Perna N T, Burland V, Riley M, Collado-Vides J, Glasner J D, Rode C K, Mayhew G F, et al. Science. 1997;277:1453–1474. - PubMed
-
- Tomb J F, White O, Kerlavage A R, Clayton R A, Sutton G G, Fleischmann R D, Ketchum K A, Klenk H P, Gill S, Dougherty B A, et al. Nature (London) 1997;388:539–547. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

