Escherichia coli strains lacking protein HU are UV sensitive due to a role for HU in homologous recombination
- PMID: 9683467
- PMCID: PMC107354
- DOI: 10.1128/JB.180.15.3750-3756.1998
Escherichia coli strains lacking protein HU are UV sensitive due to a role for HU in homologous recombination
Abstract
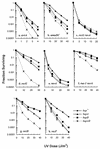
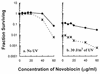
hupA and hupB encode the alpha and beta subunits of the Escherichia coli histone-like protein HU. Here we show that E. coli hup mutants are sensitive to UV in the rec+ sbc+, recBC sbcA, recBC sbcBC, umuDC, recF, and recD backgrounds. However, hupAB mutations do not enhance the UV sensitivity of resolvase-deficient recG ruvA strains. hupAB uvrA and hupAB recG strains are supersensitive to UV. hup mutations enhance the UV sensitivity of ruvA strains to a much lesser extent but enhance that of rus-1 ruvA strains to the same extent as for rus+ ruv+ strains. Our results suggest that HU plays a role in recombinational DNA repair that is not specifically limited to double-strand break repair or daughter strand gap repair; the lack of HU affects the RecG RusA and RuvABC pathways for Holliday junction processing equally if the two pathways are equally active in recombinational repair; the function of HU is not in the substrate processing step or in the RecFOR-directed synapsis action during recombinational repair. Furthermore, the UV sensitivity of hup mutants cannot be suppressed by overexpression of wild-type or mutant gyrB, which confers novobiocin resistance, or by different concentrations of a gyrase inhibitor that can increase or decrease the supercoiling of chromosomal DNA.
Figures
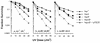
References
-
- Bensaid A, Almeida A, Drlica K, Rouviere-Yaniv J. Cross-talk between topoisomerase I and HU in Escherichia coli. J Mol Biol. 1996;256:292–300. - PubMed
-
- Bonnefoy E, Takahashi M, Rouviere-Yaniv J. Quantitative determination of DNA-binding parameters show a specific interaction of the HU protein of Escherichia coli to cruciform DNA. J Mol Biol. 1994;242:116–129. - PubMed
-
- Brendel M, Haynes R H. Interactions among genes controlling sensitivity to radiation and alkylation in yeast. Mol Gen Genet. 1973;125:197–216. - PubMed
-
- Castaing B, Zelwer C, Laval J, Boiteux S. HU protein of Escherichia coli binds specifically to DNA that contains single-strand breaks or gaps. J Biol Chem. 1995;270:10291–10296. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

