Switch from translation to RNA replication in a positive-stranded RNA virus
- PMID: 9694795
- PMCID: PMC317040
- DOI: 10.1101/gad.12.15.2293
Switch from translation to RNA replication in a positive-stranded RNA virus
Abstract
In positive-stranded viruses, the genomic RNA serves as a template for both translation and RNA replication. Using poliovirus as a model, we examined the interaction between these two processes. We show that the RNA polymerase is unable to replicate RNA templates undergoing translation. We discovered that an RNA structure at the 5' end of the viral genome, next to the internal ribosomal entry site, carries signals that control both viral translation and RNA synthesis. The interaction of this RNA structure with the cellular factor PCBP up-regulates viral translation, while the binding of the viral protein 3CD represses translation and promotes negative-strand RNA synthesis. We propose that the interaction of 3CD with this RNA structure controls whether the genomic RNA is used for translation or RNA replication.
Figures
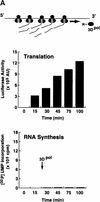

 ) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.
) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.
 ) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.
) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.
 ) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.
) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.
 ) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.
) into oocytes.
Luciferase activity was determined in oocyte extracts after 3 hr of
incubation at 22°C and expressed in AU. (D) Viral proteins
3Dpol, 3CD, and P3 copurified with the viral translation
inhibitory activity. Western blot analysis of fractions 1–14 eluted
from the HiTrap SP column is shown. Two microliters of each fraction
was resolved in a 10% SDS–polyacrylamide gel, transferred to
nitrocelluose membrane, and probed with specific anti-3CD antibodies.
The electrophoretic mobility of P3, 3CD, and 3D is indicated at
left.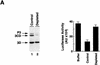
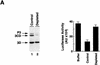
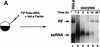
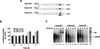
References
-
- Andino R, Rieckhof GE, Baltimore D. A functional ribonucleoprotein complex forms around the 5′ end of poliovirus RNA. Cell. 1990a;63:369–380. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
