In vitro selection and characterization of human immunodeficiency virus type 1 variants with increased resistance to ABT-378, a novel protease inhibitor
- PMID: 9696850
- PMCID: PMC109995
- DOI: 10.1128/JVI.72.9.7532-7541.1998
In vitro selection and characterization of human immunodeficiency virus type 1 variants with increased resistance to ABT-378, a novel protease inhibitor
Abstract
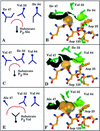
ABT-378, a new human immunodeficiency virus type 1 (HIV-1) protease inhibitor which is significantly more active than ritonavir in cell culture, is currently under investigation for the treatment of AIDS. Development of viral resistance to ABT-378 in vitro was studied by serial passage of HIV-1 (pNL4-3) in MT-4 cells. Selection of viral variants with increasing concentrations of ABT-378 revealed a sequential appearance of mutations in the protease gene: I84V-L10F-M46I-T91S-V32I-I47V. Further selection at a 3.0 microM inhibitor concentration resulted in an additional change at residue 47 (V47A), as well as reversion at residue 32 back to the wild-type sequence. The 50% effective concentration of ABT-378 against passaged virus containing these additional changes was 338-fold higher than that against wild-type virus. In addition to changes in the protease gene, sequence analysis of passaged virus revealed mutations in the p1/p6 (P1' residue Leu to Phe) and p7/p1 (P2 residue Ala to Val) gag proteolytic processing sites. The p1/p6 mutation appeared in several clones derived from early passages and was present in all clones obtained from passage P11 (0.42 microM ABT-378) onward. The p7/p1 mutation appeared very late during the selection process and was strongly associated with the emergence of the additional change at residue 47 (V47A) and the reversion at residue 32 back to the wild-type sequence. Furthermore, this p7/p1 mutation was present in all clones obtained from passage P17 (3.0 microM ABT-378) onward and always occurred in conjunction with the p1/p6 mutation. Full-length molecular clones containing protease mutations observed very late during the selection process were constructed and found to be viable only in the presence of both the p7/p1 and p1/p6 cleavage-site mutations. This suggests that mutation of these gag proteolytic cleavage sites is required for the growth of highly resistant HIV-1 selected by ABT-378 and supports recent work demonstrating that mutations in the p7/p1/p6 region play an important role in conferring resistance to protease inhibitors (L. Doyon et al., J. Virol. 70:3763-3769, 1996; Y. M. Zhang et al., J. Virol. 71:6662-6670, 1997).
Figures
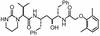




References
-
- Barrie K A, Perez E E, Lamers S L, Farmerie W G, Dunn B M, Sleasman J W, Goodenow M M. Natural variation in HIV-1 protease, Gag p7 and p6, and protease cleavage sites within gag/pol polyproteins: amino acid substitutions in the absence of protease inhibitors in mothers and children infected by human immunodeficiency virus type 1. Virology. 1996;219:407–416. - PubMed
-
- Carrillo, A., and A. Molla. Unpublished data.
-
- Chen Z, Li Y, Schock H B, Hall D, Chen E, Kuo L C. Three-dimensional structure of a mutant HIV-1 protease displaying cross-resistance to all protease inhibitors in clinical trials. J Biol Chem. 1995;270:21433–21436. - PubMed
-
- Collins J R, Burt S K, Erickson J W. Activated dynamics of flap opening in HIV-1 protease. Adv Exp Med Biol. 1995;362:455–460. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

