Complementation of the Arabidopsis pds1 mutation with the gene encoding p-hydroxyphenylpyruvate dioxygenase
- PMID: 9701587
- PMCID: PMC34895
- DOI: 10.1104/pp.117.4.1317
Complementation of the Arabidopsis pds1 mutation with the gene encoding p-hydroxyphenylpyruvate dioxygenase
Abstract
Plastoquinone and tocopherols are the two major quinone compounds in higher plant chloroplasts and are synthesized by a common pathway. In previous studies we characterized two loci in Arabidopsis defining key steps of this biosynthetic pathway. Mutation of the PDS1 locus disrupts the activity of p-hydroxyphenylpyruvate dioxygenase (HPPDase), the first committed step in the synthesis of both plastoquinone and tocopherols in plants. Although plants homozygous for the pds1 mutation could be rescued by growth in the presence of homogentisic acid, the product of HPPDase, we were unable to determine if the mutation directly or indirectly disrupted HPPDase activity. This paper reports the isolation of a cDNA, pHPPD, encoding Arabidopsis HPPDase and its functional characterization by expression in both plants and Escherichia coli. pHPPD encodes a 50-kD polypeptide with homology to previously identified HPPDases, including 37 highly conserved amino acid residues clustered in the carboxyl region of the protein. Expression of pHPPD in E. coli catalyzes the accumulation of homogentisic acid, indicating that it encodes a functional HPPDase enzyme. Mapping of pHPPD and co-segregation analysis of the pds1 mutation and the HPPD gene indicate tight linkage. Constitutive expression of pHPPD in a pds1 mutant background complements this mutation. Finally, comparison of the HPPD genomic sequences from wild type and pds1 identified a 17-bp deletion in the pds1 allele that results in deletion of the carboxyterminal 26 amino acids of the HPPDase protein. Together, these data conclusively demonstrate that pds1 is a mutation in the HPPDase structural gene.
Figures
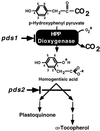


References
-
- Altschul SF, Gish W, Miller W, Myers E, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Bent A, Kunkel BN, Dahlbeck D, Brown KL, Schmidt R, Giraudat J, Leung J, Staskawicz BJ. RPS2 of Arabidopsis thaliana: a leucine-rich repeat class of plant disease resistance genes. Science. 1994;265:1856–1860. - PubMed
-
- Berger S, Ellersiek U, Westoff P, Steinmuller K. Studies on the expression of NDH-H, a subunit of the NAD(P)H-plastoquinone-oxidoreductase of higher plant chloroplasts. Planta. 1993;190:25–31.
-
- DellaPorta S, Wood J, Hicks J. A plant DNA minipreparation: version 2. Plant Mol Biol Rep. 1983;1:19–22.
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

