Polarity of DNA strand exchange promoted by recombination proteins of the RecA family
- PMID: 9707563
- PMCID: PMC21424
- DOI: 10.1073/pnas.95.17.9843
Polarity of DNA strand exchange promoted by recombination proteins of the RecA family
Abstract
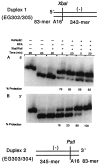
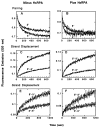
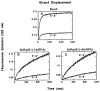
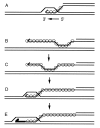
Homologs of Escherichia coli RecA recombination protein, which have been found throughout the living kingdom, promote homologous pairing and strand exchange. The nucleoprotein filament, within which strand exchange occurs, has been conserved through evolution, but conservation of the polarity of exchange and the significance of that directionality has not been settled. Using oligonucleotides as substrates, and assays based on fluorescence resonance energy transfer (FRET), we distinguished the biased formation of homologous joints at either end of duplex DNA from the subsequent directionality of strand exchange. As with E. coli RecA protein, the homologous Rad51 proteins from both Homo sapiens (HsRad51) and Saccharomyces cerevisiae (ScRad51) propagated DNA strand exchange preferentially in the 5' to 3' direction. The data suggest that 5' to 3' polarity is a conserved intrinsic property of recombination filaments.
Figures

References
-
- Shinohara A, Ogawa H, Matsuda Y, Ushio N, Ikeo K, Ogawa T. Nat Genet. 1993;4:239–243. - PubMed
-
- Kowalczykowski S C, Eggleston A K. Annu Rev Biochem. 1994;63:991–1043. - PubMed
-
- Shinohara A, Ogawa H, Ogawa T. Cell. 1992;69:457–470. - PubMed
-
- Bishop D K, Park D, Xu L, Kleckner N. Cell. 1992;69:439–456. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

