SirR, a novel iron-dependent repressor in Staphylococcus epidermidis
- PMID: 9712757
- PMCID: PMC108495
- DOI: 10.1128/IAI.66.9.4123-4129.1998
SirR, a novel iron-dependent repressor in Staphylococcus epidermidis
Abstract
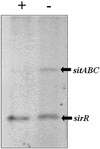
In Staphylococcus epidermidis and Staphylococcus aureus, a number of cell wall- and cytoplasmic membrane-associated lipoproteins are induced in response to iron starvation. To gain insights into the molecular basis of iron-dependent gene regulation in the staphylococci, we sequenced the DNA upstream of the 3-kb S. epidermidis sitABC operon, which Northern blot analysis indicates is transcriptionally regulated by the growth medium iron content. We identified two DNA sequences which are homologous to elements of the Corynebacterium diphtheriae DtxR regulon, which controls, in response to iron stress, for example, production of diphtheria toxin, siderophore, and a heme oxygenase. Upstream of the sitABC operon and divergently transcribed lies a 645-bp open reading frame (ORF), which codes for a polypeptide of approximately 25 kDa with homology to the DtxR family of metal-dependent repressor proteins. This ORF has been designated SirR (staphylococcal iron regulator repressor). Within the sitABC promoter/operator region, we also located a region of dyad symmetry overlapping the transcriptional start of sitABC which shows high homology to the DtxR operator consensus sequence, suggesting that this region, termed the Sir box, is the SirR-binding site. The SirR protein was overexpressed, purified, and used in DNA mobility shift assays; SirR retarded the migration of a synthetic oligonucleotide based on the Sir box in a metal (Fe2+ or Mn2+)-dependent manner, providing confirmatory evidence that this motif is the SirR-binding site. Furthermore, Southern blot analysis of staphylococcal chromosomal DNA with the synthetic Sir box as a probe confirmed that there are at least five Sir boxes in the S. epidermidis genome and at least three in the genome of S. aureus, suggesting that SirR controls the expression of multiple target genes. Using a monospecific polyclonal antibody raised against SirR to probe Western blots of whole-cell lysates of S. aureus, S. carnosus, S. epidermidis, S. hominis, S. cohnii, S. lugdunensis, and S. haemolyticus, we identified an approximately 25-kDa cross-reactive protein in each of the staphylococcal species examined. Taken together, these data suggest that SirR functions as a divalent metal cation-dependent transcriptional repressor which is widespread among the staphylococci.
Figures


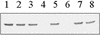



References
-
- Arbuthnott J P, Arbuthnott E, Arbuthnott A D J, Pike W J, Cockayne A. Investigation of microbial growth in vivo: evaluation of a novel in vivo chamber implant system. FEMS Microbiol Lett. 1992;100:75–80. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. New York, N.Y: Greene Publishing Associates and Wiley-Interscience; 1994.
-
- Bsat N, Helmann J D. Abstracts of the 97th General Meeting of the American Society for Microbiology 1997. Washington, D.C: American Society for Microbiology; 1997. Regulation of Bacillus subtilis iron uptake genes by Fur protein, abstr. H-193.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

