Cdc34 and the F-box protein Met30 are required for degradation of the Cdk-inhibitory kinase Swe1
- PMID: 9716410
- PMCID: PMC317080
- DOI: 10.1101/gad.12.16.2587
Cdc34 and the F-box protein Met30 are required for degradation of the Cdk-inhibitory kinase Swe1
Abstract
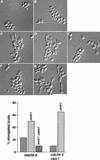
Ubiquitin-mediated proteolysis controls the abundance of many cell cycle regulatory proteins. Recent work in Saccharomyces cerevisiae suggests that a complex consisting of Cdc53, Skp1, and a third component known as an F-box protein (termed SCF) in combination with Cdc34 specifically targets regulatory proteins for degradation, and that substrate specificity is likely to be mediated by the F-box subunit. A screen for genetic interactions with a cdc34 mutation yielded MET30, which encodes an F-box protein. MET30 is an essential gene required for cell cycle progression and met30 mutations interact genetically with mutations in SCF components. Furthermore, physical interactions between Met30, Cdc53, Cdc34, and Skp1 in vivo provide evidence for an SCFMet30 complex. We demonstrate the involvement of Met30 in the degradation of the Cdk-inhibitory kinase Swe1. Swe1 is stabilized in met30 mutants and GST-Met30 pull-down experiments reveal that Met30 specifically binds Swe1 in vivo. Furthermore, extracts prepared from cdc34 or met30 mutants are defective in polyubiquitination of Swe1. Taken together, these data suggest that SCF-mediated proteolysis may contribute to the regulation of entry into mitosis. Our data, in combination with previously published results, also provide evidence for distinct SCF complexes in vivo and support the idea that their F-box subunits mediate SCF substrate specificity.
Figures


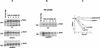

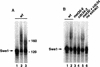


Similar articles
-
Cdc53 is a scaffold protein for multiple Cdc34/Skp1/F-box proteincomplexes that regulate cell division and methionine biosynthesis in yeast.Genes Dev. 1998 Mar 1;12(5):692-705. doi: 10.1101/gad.12.5.692. Genes Dev. 1998. PMID: 9499404 Free PMC article.
-
Regulation of transcription by ubiquitination without proteolysis: Cdc34/SCF(Met30)-mediated inactivation of the transcription factor Met4.Cell. 2000 Aug 4;102(3):303-14. doi: 10.1016/s0092-8674(00)00036-2. Cell. 2000. PMID: 10975521
-
Cdc53/cullin and the essential Hrt1 RING-H2 subunit of SCF define a ubiquitin ligase module that activates the E2 enzyme Cdc34.Genes Dev. 1999 Jun 15;13(12):1614-26. doi: 10.1101/gad.13.12.1614. Genes Dev. 1999. PMID: 10385629 Free PMC article.
-
Proteolysis and the G1-S transition: the SCF connection.Curr Opin Genet Dev. 1998 Feb;8(1):36-42. doi: 10.1016/s0959-437x(98)80059-2. Curr Opin Genet Dev. 1998. PMID: 9529603 Review.
-
Cell cycle regulation by the ubiquitin pathway.FASEB J. 1997 Nov;11(13):1067-75. doi: 10.1096/fasebj.11.13.9367342. FASEB J. 1997. PMID: 9367342 Review.
Cited by
-
Studying interactions of four proteins in the yeast two-hybrid system: structural resemblance of the pVHL/elongin BC/hCUL-2 complex with the ubiquitin ligase complex SKP1/cullin/F-box protein.Proc Natl Acad Sci U S A. 1999 Aug 17;96(17):9533-8. doi: 10.1073/pnas.96.17.9533. Proc Natl Acad Sci U S A. 1999. PMID: 10449727 Free PMC article.
-
SCF ubiquitin protein ligases and phosphorylation-dependent proteolysis.Philos Trans R Soc Lond B Biol Sci. 1999 Sep 29;354(1389):1533-50. doi: 10.1098/rstb.1999.0497. Philos Trans R Soc Lond B Biol Sci. 1999. PMID: 10582239 Free PMC article.
-
The yeast ubiquitin ligase SCFMet30: connecting environmental and intracellular conditions to cell division.Cell Div. 2006 Aug 8;1:16. doi: 10.1186/1747-1028-1-16. Cell Div. 2006. PMID: 16895602 Free PMC article.
-
Prediction of yeast protein-protein interaction network: insights from the Gene Ontology and annotations.Nucleic Acids Res. 2006 Apr 26;34(7):2137-50. doi: 10.1093/nar/gkl219. Print 2006. Nucleic Acids Res. 2006. PMID: 16641319 Free PMC article.
-
Protein arginine methyltransferases: from unicellular eukaryotes to humans.Eukaryot Cell. 2007 Jun;6(6):889-98. doi: 10.1128/EC.00099-07. Epub 2007 Apr 27. Eukaryot Cell. 2007. PMID: 17468392 Free PMC article. Review. No abstract available.
References
-
- Aligue R, Wu L, Russell P. Regulation of Schizosaccharomyces pombe Wee1 tyrosine kinase. J Biol Chem. 1997;272:13320–13325. - PubMed
-
- Amon A, Surana U, Muroff I, Nasmyth K. Regulation of p34CDC28 tyrosine phosphorylation is not required for entry into mitosis in S. cerevisiae. Nature. 1992;355:368–371. - PubMed
-
- Bai C, Sen P, Hofman K, Ma L, Goebl M, Harper JW, Elledge SJ. SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box. Cell. 1996;86:263–274. - PubMed
-
- Barral Y, Jentsch S, Mann C. G1 cyclin turnover and nutrient uptake are controlled by a common pathway in yeast. Genes & Dev. 1995;9:399–409. - PubMed
-
- Beers EP, Callis J. Utility of polyhistidine-tagged ubiquitin in the purification of ubiquitin-protein conjugates and as an affinity ligand for the purification of ubiquitin-specific hydrolases. J Biol Chem. 1993;268:21645–21649. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
