Purification and characterization of peroxidases correlated with lignification in poplar xylem
- PMID: 9733532
- PMCID: PMC34849
- DOI: 10.1104/pp.118.1.125
Purification and characterization of peroxidases correlated with lignification in poplar xylem
Abstract
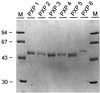
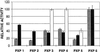
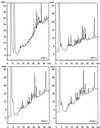
Lignin is an integral cell wall component of all vascular plants. Peroxidases are widely believed to catalyze the last enzymatic step in the biosynthesis of lignin, the dehydrogenation of the p-coumaryl alcohols. As the first stage in identifying lignin-specific peroxidase isoenzymes, the classical anionic peroxidases found in the xylem of poplar (Populus trichocarpa Trichobel) were purified and characterized. Five different poplar xylem peroxidases (PXP 1, PXP 2, PXP 3-4, PXP 5, and PXP 6) were isolated. All five peroxidases were strongly glycosylated (3.6% to 4.9% N-glucosamine), with apparent molecular masses between 46 and 54 kD and pI values between pH 3.1 and 3.8. Two of the five isolated peroxidases (PXP 3-4 and PXP 5) could oxidize the lignin monomer analog syringaldazine, an activity previously correlated with lignification in poplar. Because these isoenzymes were specifically or preferentially expressed in xylem, PXP 3-4 and PXP 5 are suggested to be involved in lignin polymerization.
Figures





Similar articles
-
The syringaldazine-oxidizing peroxidase PXP 3-4 from poplar xylem: cDNA isolation, characterization and expression.Plant Mol Biol. 2001 Nov;47(5):581-93. doi: 10.1023/a:1012271729285. Plant Mol Biol. 2001. PMID: 11725944
-
Characterization of basic p-coumaryl and coniferyl alcohol oxidizing peroxidases from a lignin-forming Picea abies suspension culture.Plant Mol Biol. 2005 May;58(2):141-57. doi: 10.1007/s11103-005-5345-6. Plant Mol Biol. 2005. PMID: 16027971
-
The role of xylem class III peroxidases in lignification.J Exp Bot. 2009;60(2):367-76. doi: 10.1093/jxb/ern278. J Exp Bot. 2009. PMID: 19264758 Review.
-
Monolignol oxidation by xylem peroxidase isoforms of Norway spruce (Picea abies) and silver birch (Betula pendula).Tree Physiol. 2006 May;26(5):605-11. doi: 10.1093/treephys/26.5.605. Tree Physiol. 2006. PMID: 16452074
-
Cell wall lignin is polymerised by class III secretable plant peroxidases in Norway spruce.J Integr Plant Biol. 2010 Feb;52(2):186-94. doi: 10.1111/j.1744-7909.2010.00928.x. J Integr Plant Biol. 2010. PMID: 20377680 Review.
Cited by
-
Apoplast proteome reveals that extracellular matrix contributes to multistress response in poplar.BMC Genomics. 2010 Nov 29;11:674. doi: 10.1186/1471-2164-11-674. BMC Genomics. 2010. PMID: 21114852 Free PMC article.
-
RNA interference identifies a calcium-dependent protein kinase involved in Medicago truncatula root development.Plant Cell. 2005 Nov;17(11):2911-21. doi: 10.1105/tpc.105.035394. Epub 2005 Sep 30. Plant Cell. 2005. PMID: 16199614 Free PMC article.
-
Chorion peroxidase-mediated NADH/O(2) oxidoreduction cooperated by chorion malate dehydrogenase-catalyzed NADH production: a feasible pathway leading to H(2)O(2) formation during chorion hardening in Aedes aegypti mosquitoes.Biochim Biophys Acta. 2000 Oct 18;1523(2-3):246-53. doi: 10.1016/s0304-4165(00)00129-x. Biochim Biophys Acta. 2000. PMID: 11042391 Free PMC article.
-
Enhancing digestibility and ethanol yield of Populus wood via expression of an engineered monolignol 4-O-methyltransferase.Nat Commun. 2016 Jun 28;7:11989. doi: 10.1038/ncomms11989. Nat Commun. 2016. PMID: 27349324 Free PMC article.
-
Autotoxicity mechanism of Oryza sativa: transcriptome response in rice roots exposed to ferulic acid.BMC Genomics. 2013 May 25;14:351. doi: 10.1186/1471-2164-14-351. BMC Genomics. 2013. PMID: 23705659 Free PMC article.
References
-
- Baier M, Goldberg R, Catesson AM, Francesch C, Rolando C. Seasonal changes of isoperoxidases from poplar bark tissues. Phytochemistry. 1993;32:789–793.
-
- Barkholt V, Jensen AL. Amino acid analysis: determination of cysteine plus half-cystine in proteins after hydrochloric acid hydrolysis with a disulfide compound as additive. Anal Biochem. 1989;177:318–322. - PubMed
-
- Boerjan W, Baucher M, Chabbert B, Petit-Conil M, Leplé J-C, Pilate G, Cornu D, Monties B, Inzé D, Van Doorsselaere J, and others (1997) Genetic modification of lignin biosynthesis in quaking aspen (Populus tremuloides) and poplar (Populus tremula × Populus alba). In NB Klopfenstein, YW Chun, M-S Kim, MR Ahuja, eds, Micropropagation, Genetic Engineering, and Molecular Biology of Populus (General Technical Report RM-GTR-297). Rocky Mountain Forest and Range Experiment Station, Fort Collins, CO, pp 193–205
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

