Oxidative stress response and characterization of the oxyR-ahpC and furA-katG loci in Mycobacterium marinum
- PMID: 9733688
- PMCID: PMC107510
- DOI: 10.1128/JB.180.18.4856-4864.1998
Oxidative stress response and characterization of the oxyR-ahpC and furA-katG loci in Mycobacterium marinum
Abstract
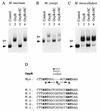
Oxidative stress response in pathogenic mycobacteria is believed to be of significance for host-pathogen interactions at various stages of infection. It also plays a role in determining the intrinsic susceptibility to isoniazid in mycobacterial species. In this work, we characterized the oxyR-ahpC and furA-katG loci in the nontuberculous pathogen Mycobacterium marinum. In contrast to Mycobacterium smegmatis and like Mycobacterium tuberculosis and Mycobacterium leprae, M. marinum was shown to possess a closely linked and divergently oriented equivalents of the regulator of peroxide stress response oxyR and its subordinate gene ahpC, encoding a homolog of alkyl hydroperoxide reductase. Purified mycobacterial OxyR was found to bind to the oxyR-ahpC promoter region from M. marinum and additional mycobacterial species. Mobility shift DNA binding analyses using OxyR binding sites from several mycobacteria and a panel of in vitro-generated mutants validated the proposed consensus mycobacterial recognition sequence. M. marinum AhpC levels detected by immunoblotting, were increased upon treatment with H2O2, in keeping with the presence of a functional OxyR and its binding site within the promoter region of ahpC. In contrast, OxyR did not bind to the sequences upstream of the katG structural gene, and katG expression did not follow the pattern seen with ahpC. Instead, a new open reading frame encoding a homolog of the ferric uptake regulator Fur was identified immediately upstream of katG in M. marinum. The furA-katG linkage and arrangement are ubiquitous in mycobacteria, suggesting the presence of additional regulators of oxidative stress response and potentially explaining the observed differences in ahpC and katG expression. Collectively, these findings broaden our understanding of oxidative stress response in mycobacteria. They also suggest that M. marinum will be useful as a model system for studying the role of oxidative stress response in mycobacterial physiology, intracellular survival, and other host-pathogen interactions associated with mycobacterial diseases.
Figures


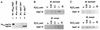


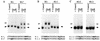


References
-
- Altuvia S, Almirom M, Huisman G, Kolter R, Storz G. The dps promoter is activated by OxyR during growth and by IHF and ςS in stationary phase. Mol Microbiol. 1994;13:265–272. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons; 1989.
-
- Chae H Z, Robinson K, Poole L B, Church G, Storz G, Rhee S G. Cloning and sequencing of thiol-specific antioxidant from mammalian brain: alkyl hydroperoxide reductase and thiol-specific antioxidant define a large family of antioxidant enzymes. Proc Natl Acad Sci USA. 1994;91:7017–7021. - PMC - PubMed
-
- Chan J, Kaufmann H E. Immune mechanisms of protection. In: Bloom B R, editor. Tuberculosis: pathogenesis, protection, and control. Washington, D.C: ASM Press; 1994. pp. 389–415.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

