Activation of the Epstein-Barr virus transcription factor BZLF1 by 12-O-tetradecanoylphorbol-13-acetate-induced phosphorylation
- PMID: 9733851
- PMCID: PMC110148
- DOI: 10.1128/JVI.72.10.8105-8114.1998
Activation of the Epstein-Barr virus transcription factor BZLF1 by 12-O-tetradecanoylphorbol-13-acetate-induced phosphorylation
Abstract
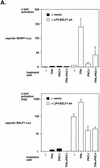
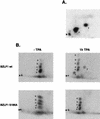
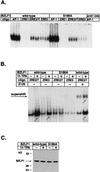
BZLF1 is a member of the extended AP-1 family of transcription factors which binds to specific BZLF1 sequence motifs within early Epstein-Barr virus (EBV) promoters and to closely related AP-1 motifs. BZLF1's activity is regulated at the transcriptional level as well as through protein interactions and posttranslational modifications. Phorbol esters or immunoglobulin cross-linking both reactivate EBV from latently infected B cells via transactivation of BZLF1. We report here that the phorbol ester 12-O-tetradecanoylphorbol-13-acetate (TPA) is capable of inducing BZLF1's activity even further. The induction occurs at the posttranscriptional level and depends on a single serine residue located in the DNA binding domain of BZLF1. This serine residue (S186) is phosphorylated by protein kinase C in vitro and in vivo after stimulation with TPA. Phosphorylation of S186 per se interferes with the DNA binding affinity of BZLF1 in vitro but is mandatory for TPA-induced increase in DNA binding of BZLF1, as shown in gel retardation assays and reconstruction experiments with cellular extracts. In transcriptional reporter assays, S186 is essential for the activation of BZLF1 by TPA. Presumably, a yet-to-be-identified cellular factor restores the DNA binding affinity and enhances the transcriptional activity of S186-phosphorylated BZLF1, which is required to induce the lytic phase of EBV's life cycle.
Figures






References
-
- Aderem A. The MARCKS brothers: a family of protein kinase C substrates. Cell. 1992;71:713–716. - PubMed
-
- Azzi A, Boscoboinik D, Hensey C. The protein kinase C family. Eur J Biochem. 1992;208:547–557. - PubMed
-
- Baer R, Bankier A T, Biggin M D, Deininger P L, Farrell P J, Gibson T J, Hatfull G, Hudson G S, Satchwell S C, Seguin C, Tufnell P S, Barell B G. DNA sequence and expression of the B95-8 Epstein-Barr virus genome. Nature (London) 1984;310:207–211. - PubMed
-
- Binetruy B, Smeal T, Karin M. Ha-Ras augments c-Jun activity and stimulates phosphorylation of its activation domain. Nature (London) 1991;351:122–127. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

