Indirect mutagenesis by oxidative DNA damage: formation of the pyrimidopurinone adduct of deoxyguanosine by base propenal
- PMID: 9736698
- PMCID: PMC21604
- DOI: 10.1073/pnas.95.19.11113
Indirect mutagenesis by oxidative DNA damage: formation of the pyrimidopurinone adduct of deoxyguanosine by base propenal
Abstract
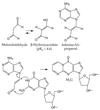
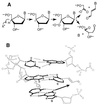
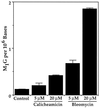
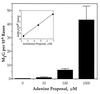
Oxidation of endogenous macromolecules can generate electrophiles capable of forming mutagenic adducts in DNA. The lipid peroxidation product malondialdehyde, for example, reacts with DNA to form M1G, the mutagenic pyrimidopurinone adduct of deoxyguanosine. In addition to free radical attack of lipids, DNA is also continuously subjected to oxidative damage. Among the products of oxidative DNA damage are base propenals. We hypothesized that these structural analogs of malondialdehyde would react with DNA to form M1G. Consistent with this hypothesis, we detected a dose-dependent increase in M1G in DNA treated with calicheamicin and bleomycin, oxidizing agents known to produce base propenal. The hypothesis was proven when we determined that 9-(3-oxoprop-1-enyl)adenine gives rise to the M1G adduct with greater efficiency than malondialdehyde itself. The reactivity of base propenals to form M1G and their presence in the target DNA suggest that base propenals derived from oxidative DNA damage may contribute to the mutagenic burden of a cell.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

