Association of guide RNA binding protein gBP21 with active RNA editing complexes in Trypanosoma brucei
- PMID: 9742118
- PMCID: PMC109187
- DOI: 10.1128/MCB.18.10.6014
Association of guide RNA binding protein gBP21 with active RNA editing complexes in Trypanosoma brucei
Abstract
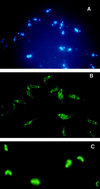
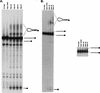
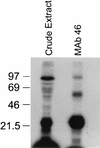
RNA editing in Trypanosoma brucei mitochondria produces mature mRNAs by a series of enzyme-catalyzed reactions that specifically insert or delete uridylates in association with a macromolecular complex. Using a mitochondrial fraction enriched for in vitro RNA editing activity, we produced several monoclonal antibodies that are specific for a 21-kDa guide RNA (gRNA) binding protein initially identified by UV cross-linking. Immunofluorescence studies localize the protein to the mitochondrion, with a preference for the kinetoplast. The antibodies cause a supershift of previously identified gRNA-specific ribonucleoprotein complexes and immunoprecipitate in vitro RNA editing activities that insert and delete uridylates. The immunoprecipitated material also contains gRNA-specific endoribonuclease, terminal uridylyltransferase, and RNA ligase activities as well as gRNA and both edited and unedited mRNA. The immunoprecipitate contains numerous proteins, of which the 21-kDa protein, a 90-kDa protein, and novel 55- and 16-kDa proteins can be UV cross-linked to gRNA. These studies indicate that the 21-kDa protein associates with the ribonucleoprotein complex (or complexes) that catalyze RNA editing.
Figures

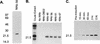

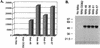
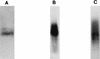

References
-
- Benne R. RNA editing in trypanosomes. Eur J Biochem. 1994;221:9–23. - PubMed
-
- Benne R, van den Burg J, Brakenhoff J P J, Sloof P, Van Boom J H, Tromp M C. Major transcripts of the frameshifted coxII gene from trypanosome mitochondria contain four nucleotides that are not encoded in the DNA. Cell. 1986;46:819–826. - PubMed
-
- Bhat G J, Koslowsky D J, Feagin J E, Smiley B L, Stuart K. An extensively edited mitochondrial transcript in kinetoplastids encodes a protein homologous to ATPase subunit 6. Cell. 1990;61:885–894. - PubMed
-
- Blum B, Bakalara N, Simpson L. A model for RNA editing in kinetoplastid mitochondria: “guide” RNA molecules transcribed from maxicircle DNA provide the edited information. Cell. 1990;60:189–198. - PubMed
-
- Blum B, Simpson L. Guide RNAs in kinetoplastid mitochondria have a nonencoded 3′ oligo(U) tail involved in recognition of the preedited region. Cell. 1990;62:391–397. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
