ClpB in a cyanobacterium: predicted structure, phylogenetic relationships, and regulation by light and temperature
- PMID: 9748452
- PMCID: PMC107555
- DOI: 10.1128/JB.180.19.5173-5182.1998
ClpB in a cyanobacterium: predicted structure, phylogenetic relationships, and regulation by light and temperature
Abstract
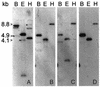
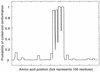
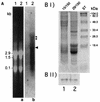
The sequence of a genomic clone encoding a 100-kDa stress protein of Plectonema boryanum (p-ClpB) was determined. The predicted polypeptide contains two putative ATPase regions located within two highly conserved domains (N1 and N2), a spacer region that likely forms a coiled-coil domain, and a highly conserved consensus CK2 phosphorylation domain. The coiled-coil region and the putative site of phosphorylation are not unique to p-ClpB; they are present in all ClpB sequences examined and are absent from the ClpB paralogs ClpA, ClpC, ClpX, and ClpY. Small quantities of a 4.5-kb p-clpB transcript and 110-kDa cytosolic p-ClpB protein were detected in cells grown under optimal conditions; however, increases in the quantities of the transcript and protein were observed in cells grown under excess light and low temperature conditions. Finally, we analyzed ClpA, ClpB, and ClpC sequences from 27 organisms in order to predict phylogenetic relationships among the homologs. We have used this information, along with an identity alignment, to redefine the Clp subfamilies.
Figures


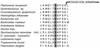

Similar articles
-
The cyanobacterium Synechococcus sp. PCC 7942 possesses a close homologue to the chloroplast ClpC protein of higher plants.Plant Mol Biol. 1996 Jul;31(4):721-30. doi: 10.1007/BF00019460. Plant Mol Biol. 1996. PMID: 8806403
-
Characterization of a chaperone ClpB homologue of Paracoccidioides brasiliensis.Yeast. 2002 Aug;19(11):963-72. doi: 10.1002/yea.888. Yeast. 2002. PMID: 12125053
-
The heat shock protein ClpB mediates the development of thermotolerance in the cyanobacterium Synechococcus sp. strain PCC 7942.J Bacteriol. 1996 Aug;178(16):4839-46. doi: 10.1128/jb.178.16.4839-4846.1996. J Bacteriol. 1996. PMID: 8759846 Free PMC article.
-
Analysis of the AAA+ chaperone clpB gene and stress-response expression in the halophilic methanogenic archaeon Methanohalophilus portucalensis.Microbiology (Reading). 2007 Aug;153(Pt 8):2572-2583. doi: 10.1099/mic.0.2007/007633-0. Microbiology (Reading). 2007. PMID: 17660421
-
The heat-shock protein ClpB in Escherichia coli is a protein-activated ATPase.J Biol Chem. 1992 Oct 5;267(28):20429-34. J Biol Chem. 1992. PMID: 1400361
Cited by
-
Structure and function of the middle domain of ClpB from Escherichia coli.Biochemistry. 2003 Dec 9;42(48):14242-8. doi: 10.1021/bi035573d. Biochemistry. 2003. PMID: 14640692 Free PMC article.
-
Low-temperature-induced accumulation of xanthophylls and its structural consequences in the photosynthetic membranes of the cyanobacterium Cylindrospermopsis raciborskii: an FTIR spectroscopic study.Proc Natl Acad Sci U S A. 2002 Feb 19;99(4):2410-5. doi: 10.1073/pnas.042698799. Epub 2002 Feb 12. Proc Natl Acad Sci U S A. 2002. PMID: 11842219 Free PMC article.
-
ClpB1 overproduction in Synechocystis sp. strain PCC 6803 increases tolerance to rapid heat shock.Appl Environ Microbiol. 2013 Oct;79(20):6220-7. doi: 10.1128/AEM.01661-13. Epub 2013 Aug 2. Appl Environ Microbiol. 2013. PMID: 23913426 Free PMC article.
-
Protection and Damage Repair Mechanisms Contributed To the Survival of Chroococcidiopsis sp. Exposed To a Mars-Like Near Space Environment.Microbiol Spectr. 2022 Dec 21;10(6):e0344022. doi: 10.1128/spectrum.03440-22. Epub 2022 Dec 1. Microbiol Spectr. 2022. PMID: 36453906 Free PMC article.
-
Genetic analysis reveals domain interactions of Arabidopsis Hsp100/ClpB and cooperation with the small heat shock protein chaperone system.Plant Cell. 2005 Feb;17(2):559-71. doi: 10.1105/tpc.104.027540. Epub 2005 Jan 19. Plant Cell. 2005. PMID: 15659638 Free PMC article.
References
-
- Abelovich A, Kellenber D, Shilo M. Effect of photooxidative conditions on levels of superoxide dismutase in Anacystis nidulans. Photochem Photobiol. 1974;19:379–382. - PubMed
-
- Allen M. Simple conditions for the growth of unicellular blue-green algae on plates. J Phycol. 1968;4:1–3. - PubMed
-
- Ang D, Liberek K, Skowyra D, Zylicz M, Georgopolous C. Biological role and regulation of the universally conserved heat shock proteins. J Biol Chem. 1991;266:24233–24236. - PubMed
-
- Asada K, Yoshikama K, Takahashi M, Maeda Y, Enmanji K. Superoxide dismutase from a blue green alga, Plectonema boryanum. J Biol Chem. 1975;250:2801–2807. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

