Carbon-source-dependent expression of the PalkB promoter from the Pseudomonas oleovorans alkane degradation pathway
- PMID: 9748457
- PMCID: PMC107560
- DOI: 10.1128/JB.180.19.5218-5226.1998
Carbon-source-dependent expression of the PalkB promoter from the Pseudomonas oleovorans alkane degradation pathway
Abstract
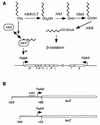
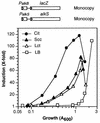
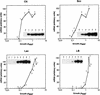
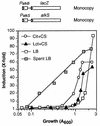
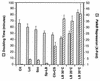
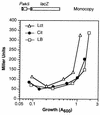
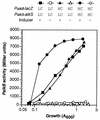
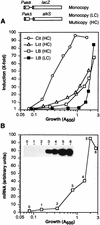
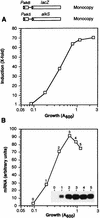
Pseudomonas oleovorans GPo1 can metabolize medium-chain-length alkanes by means of an enzymatic system whose induction is regulated by the AlkS protein. In the presence of alkanes, AlkS activates the expression of promoter PalkB, from which most of the genes of the pathway are transcribed. In addition, expression of the first enzyme of the pathway, alkane hydroxylase, is known to be influenced by the carbon source present in the growth medium, indicating the existence of an additional overimposed level of regulation associating expression of the alk genes with the metabolic status of the cell. Reporter strains bearing PalkB-lacZ transcriptional fusions were constructed to analyze the influence of the carbon source on induction of the PalkB promoter by a nonmetabolizable inducer. Expression was most efficient when cells grew at the expense of citrate, decreasing significantly when the carbon source was lactate or succinate. When cells were grown in Luria-Bertani rich medium, PalkB was strongly down-regulated. This effect was partially relieved when multiple copies of the gene coding for the AlkS activator were present and was not observed when the promoter was moved to Escherichia coli, a heterologous genetic background. Possible mechanisms responsible for PalkB regulation are discussed.
Figures
References
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons, Inc.; 1989.
-
- Bachmann B J. Derivations and genotypes of some mutant derivatives of Escherichia coli K12. In: Neidhardt F C, Ingraham J L, Low K B, Magasanik B, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella typhimurium: cellular and molecular biology. Washington, D.C: American Society for Microbiology; 1987. pp. 1191–1219.
-
- Bagdasarian M, Lurz R, Rückert B, Franklin F C H, Bagdasarian M M, Frey J, Timmis K N. Specific-purpose plasmid cloning vectors. Broad host range, high copy number, RSF1010-derived vectors, and a host-vector system for gene cloning in Pseudomonas. Gene. 1981;16:237–247. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

