Down-regulation of transmembrane carbonic anhydrases in renal cell carcinoma cell lines by wild-type von Hippel-Lindau transgenes
- PMID: 9770531
- PMCID: PMC22876
- DOI: 10.1073/pnas.95.21.12596
Down-regulation of transmembrane carbonic anhydrases in renal cell carcinoma cell lines by wild-type von Hippel-Lindau transgenes
Abstract
To discover genes involved in von Hippel-Lindau (VHL)-mediated carcinogenesis, we used renal cell carcinoma cell lines stably transfected with wild-type VHL-expressing transgenes. Large-scale RNA differential display technology applied to these cell lines identified several differentially expressed genes, including an alpha carbonic anhydrase gene, termed CA12. The deduced protein sequence was classified as a one-pass transmembrane CA possessing an apparently intact catalytic domain in the extracellular CA module. Reintroduced wild-type VHL strongly inhibited the overexpression of the CA12 gene in the parental renal cell carcinoma cell lines. Similar results were obtained with CA9, encoding another transmembrane CA with an intact catalytic domain. Although both domains of the VHL protein contribute to regulation of CA12 expression, the elongin binding domain alone could effectively regulate CA9 expression. We mapped CA12 and CA9 loci to chromosome bands 15q22 and 17q21.2 respectively, regions prone to amplification in some human cancers. Additional experiments are needed to define the role of CA IX and CA XII enzymes in the regulation of pH in the extracellular microenvironment and its potential impact on cancer cell growth.
Figures

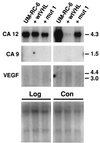
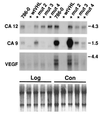
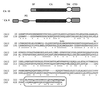


References
-
- Latif F, Tory K, Gnarra J, Yao M, Duh F M, Orcutt M L, Stackhouse T, Kuzmin I, Modi W, Geil L, et al. Science. 1993;260:1317–1320. - PubMed
-
- Kishida T, Stackhouse T M, Chen F, Lerman M I, Zbar B. Cancer Res. 1995;55:4544–4548. - PubMed
-
- Duan D R, Pause A, Burgess W H, Aso T, Chen D Y, Garrett K P, Conaway R C, Conaway J W, Linehan W M, Klausner R D. Science. 1995;269:1402–1406. - PubMed
-
- Kibel A, Iliopoulos O, DeCaprio J A, Kaelin W G., Jr Science. 1995;269:1444–1446. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

