Ectopic expression of cdc2/cdc28 kinase subunit Homo sapiens 1 uncouples cyclin B metabolism from the mitotic spindle cell cycle checkpoint
- PMID: 9774639
- PMCID: PMC109209
- DOI: 10.1128/MCB.18.11.6224
Ectopic expression of cdc2/cdc28 kinase subunit Homo sapiens 1 uncouples cyclin B metabolism from the mitotic spindle cell cycle checkpoint
Abstract
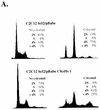
Primary human fibroblasts arrest growth in response to the inhibition of mitosis by mitotic spindle-depolymerizing drugs. We show that the mechanism of mitotic arrest is transient and implicates a decrease in the expression of cdc2/cdc28 kinase subunit Homo sapiens 1 (CKsHs1) and a delay in the metabolism of cyclin B. Primary human fibroblasts infected with a retroviral vector that drives the expression of a mutant p53 protein failed to downregulate CKsHs1 expression, degraded cyclin B despite the absence of chromosomal segregation, and underwent DNA endoreduplication. In addition, ectopic expression of CKsHs1 interfered with the control of cyclin B metabolism by the mitotic spindle cell cycle checkpoint and resulted in a higher tendency to undergo DNA endoreduplication. These results demonstrate that an altered regulation of CKsHs1 and cyclin B in cells that carry mutant p53 undermines the mitotic spindle cell cycle checkpoint and facilitates the development of aneuploidy. These data may contribute to the understanding of the origin of heteroploidy in mutant p53 cells.
Figures











Similar articles
-
p21(Waf1/Cip1) inhibition of cyclin E/Cdk2 activity prevents endoreduplication after mitotic spindle disruption.Mol Cell Biol. 1999 Jan;19(1):205-15. doi: 10.1128/MCB.19.1.205. Mol Cell Biol. 1999. PMID: 9858545 Free PMC article.
-
Protein phosphatase 2A regulates MPF activity and sister chromatid cohesion in budding yeast.Curr Biol. 1996 Dec 1;6(12):1609-20. doi: 10.1016/s0960-9822(02)70784-7. Curr Biol. 1996. PMID: 8994825
-
DNA damage during the spindle-assembly checkpoint degrades CDC25A, inhibits cyclin-CDC2 complexes, and reverses cells to interphase.Mol Biol Cell. 2003 Oct;14(10):3989-4002. doi: 10.1091/mbc.e03-03-0168. Epub 2003 Jul 25. Mol Biol Cell. 2003. PMID: 14517313 Free PMC article.
-
Gain of function properties of mutant p53 proteins at the mitotic spindle cell cycle checkpoint.Histol Histopathol. 2000 Apr;15(2):551-6. doi: 10.14670/HH-15.551. Histol Histopathol. 2000. PMID: 10809376 Review.
-
[Cyclins A and B: redundancy and specificity].Pathol Biol (Paris). 1993 Jun;41(6):547-53. Pathol Biol (Paris). 1993. PMID: 8247635 Review. French.
Cited by
-
Akt1/PKB upregulation leads to vascular smooth muscle cell hypertrophy and polyploidization.J Clin Invest. 2000 Oct;106(8):1011-20. doi: 10.1172/JCI8252. J Clin Invest. 2000. PMID: 11032861 Free PMC article.
-
Loss of cks1 homeostasis deregulates cell division cycle.J Cell Mol Med. 2010 Jan;14(1-2):154-64. doi: 10.1111/j.1582-4934.2009.00698.x. Epub 2009 Feb 17. J Cell Mol Med. 2010. PMID: 19228269 Free PMC article. Review.
-
Bridge-Induced Translocation between NUP145 and TOP2 Yeast Genes Models the Genetic Fusion between the Human Orthologs Associated With Acute Myeloid Leukemia.Front Oncol. 2017 Sep 29;7:231. doi: 10.3389/fonc.2017.00231. eCollection 2017. Front Oncol. 2017. PMID: 29034209 Free PMC article.
-
Roles of reactive oxygen species, mitochondrial membrane potential, and p53 in evodiamine-induced apoptosis and G2/M arrest of human anaplastic thyroid carcinoma cells.Chin Med. 2021 Dec 9;16(1):134. doi: 10.1186/s13020-021-00505-3. Chin Med. 2021. PMID: 34886886 Free PMC article.
References
-
- Agapova L S, Ilyinskaya G V, Turovets N A, Ivanov A V, Chumakov P M, Kopnin B P. Chromosome changes caused by alterations of p53 expression. Mutat Res. 1996;354:129–138. - PubMed
-
- Basi G, Draetta G. The cdc2 kinase: structure, activation, and its role at mitosis in vertebrate cells. In: Hutchison C, Glover D M, editors. Cell cycle control. Vol. 10. Oxford, United Kingdom: Oxford University Press; 1995. pp. 106–134.
-
- Bischoff F Z, Yim S O, Pathak S, Grant G, Siciliano M J, Giovanella B C, Strong L C, Tainsky M A. Spontaneous abnormalities in normal fibroblasts from patients with Li-Fraumeni cancer syndrome: aneuploidy and immortalization. Cancer Res. 1990;50:7979–7984. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
