Determinants of coactivator LXXLL motif specificity in nuclear receptor transcriptional activation
- PMID: 9808623
- PMCID: PMC317227
- DOI: 10.1101/gad.12.21.3357
Determinants of coactivator LXXLL motif specificity in nuclear receptor transcriptional activation
Abstract
Ligand-dependent activation of gene transcription by nuclear receptors is dependent on the recruitment of coactivators, including a family of related NCoA/SRC factors, via a region containing three helical domains sharing an LXXLL core consensus sequence, referred to as LXDs. In this manuscript, we report receptor-specific differential utilization of LXXLL-containing motifs of the NCoA-1/SRC-1 coactivator. Whereas a single LXD is sufficient for activation by the estrogen receptor, different combinations of two, appropriately spaced, LXDs are required for actions of the thyroid hormone, retinoic acid, peroxisome proliferator-activated, or progesterone receptors. The specificity of LXD usage in the cell appears to be dictated, at least in part, by specific amino acids carboxy-terminal to the core LXXLL motif that may make differential contacts with helices 1 and 3 (or 3') in receptor ligand-binding domains. Intriguingly, distinct carboxy-terminal amino acids are required for PPARgamma activation in response to different ligands. Related LXXLL-containing motifs in NCoA-1/SRC-1 are also required for a functional interaction with CBP, potentially interacting with a hydrophobic binding pocket. Together, these data suggest that the LXXLL-containing motifs have evolved to serve overlapping roles that are likely to permit both receptor-specific and ligand-specific assembly of a coactivator complex, and that these recognition motifs underlie the recruitment of coactivator complexes required for nuclear receptor function.
Figures

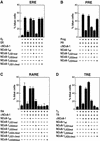






References
-
- Alland L, Muhle R, Hou H, Potes J, Chin L, Schreiber-Agus N, DePinho RA. Role for NCoR and histone deacetylase in Sin3-mediated transcriptional repression. Nature. 1997;387:49–55. - PubMed
-
- Anzick SL, Kononen J, Walker RL, Azorsa DO, Tanner MM, Guan XY, Sauter G, Kallioniemi OP, Trent JM, Meltzer PS. AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science. 1997;277:965–968. - PubMed
-
- Beekman JM, Allan GF, Tsai SY, O’Malley BW. Transcriptional activation by the estrogen receptor requires a conformational change in the ligand binding domain. Mol Endocrinol. 1993;7:1266–1274. - PubMed
-
- Bourguet W, Ruff M, Chambon P, Gronemeyer H, Moras D. Crystal structure of the ligand-binding domain of the human nuclear receptor RXR-α. Nature. 1995;375:377–382. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
