Identification of conserved, RpoS-dependent stationary-phase genes of Escherichia coli
- PMID: 9829938
- PMCID: PMC107714
- DOI: 10.1128/JB.180.23.6283-6291.1998
Identification of conserved, RpoS-dependent stationary-phase genes of Escherichia coli
Abstract
During entry into stationary phase, many free-living, gram-negative bacteria express genes that impart cellular resistance to environmental stresses, such as oxidative stress and osmotic stress. Many genes that are required for stationary-phase adaptation are controlled by RpoS, a conserved alternative sigma factor, whose expression is, in turn, controlled by many factors. To better understand the numbers and types of genes dependent upon RpoS, we employed a genetic screen to isolate more than 100 independent RpoS-dependent gene fusions from a bank of several thousand mutants harboring random, independent promoter-lacZ operon fusion mutations. Dependence on RpoS varied from 2-fold to over 100-fold. The expression of all fusion mutations was normal in an rpoS/rpoS+ merodiploid (rpoS background transformed with an rpoS-containing plasmid). Surprisingly, the expression of many RpoS-dependent genes was growth phase dependent, albeit at lower levels, even in an rpoS background, suggesting that other growth-phase-dependent regulatory mechanisms, in addition to RpoS, may control postexponential gene expression. These results are consistent with the idea that many growth-phase-regulated functions in Escherichia coli do not require RpoS for expression. The identities of the 10 most highly RpoS-dependent fusions identified in this study were determined by DNA sequence analysis. Three of the mutations mapped to otsA, katE, ecnB, and osmY-genes that have been previously shown by others to be highly RpoS dependent. The six remaining highly-RpoS-dependent fusion mutations were located in other genes, namely, gabP, yhiUV, o371, o381, f186, and o215.
Figures
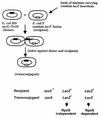



 , other RpoS-dependent genes (see text); □, genes not known to require RpoS for expression.
, other RpoS-dependent genes (see text); □, genes not known to require RpoS for expression.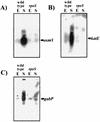

References
-
- Altschul S F, Gish W, Miller W, Myers E W, Lipman D J. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Altuvia S, Almiron M, Huisman G, Kolter R, Storz G. The dps promoter is activated by OxyR during growth and by IHF and sigma S in stationary phase. Mol Microbiol. 1994;13:265–272. - PubMed
-
- Altuvia S, Weinstein-Fischer D, Zhang A, Postow L, Storz G. A small, stable RNA induced by oxidative stress: role as a pleiotropic regulator and antimutator. Cell. 1997;90:43–53. - PubMed
-
- Bachellier S, Gilson E, Hofnung M, Hill C W. Repeated sequences. In: Neidhardt F C, Curtiss III R, Ingraham J L, Lin E C C, Low K B, Magasanik B, Reznikoff W S, Riley M, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella: cellular and molecular biology. 2nd ed. Vol. 2. Washington, D.C: ASM Press; 1996. pp. 2012–2040.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

