New small, acid-soluble proteins unique to spores of Bacillus subtilis: identification of the coding genes and regulation and function of two of these genes
- PMID: 9852018
- PMCID: PMC107777
- DOI: 10.1128/JB.180.24.6704-6712.1998
New small, acid-soluble proteins unique to spores of Bacillus subtilis: identification of the coding genes and regulation and function of two of these genes
Abstract
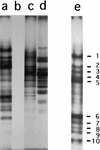
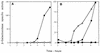
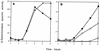
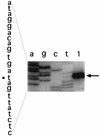
Eleven small, acid-soluble proteins (SASP) which are present in spores but not in growing cells of Bacillus subtilis were identified by sequence analysis of proteins separated by acrylamide gel electrophoresis of acid extracts from spores which lack the three major SASP (alpha, beta, and gamma). Six of these proteins are encoded by open reading frames identified previously or by analysis of the complete sequence of the B. subtilis genome, including two minor alpha/beta-type SASP (SspC and SspD) and a putative spore coat protein (CotK). Five proteins are encoded by short open reading frames that were not identified as coding regions in the analysis of the complete B. subtilis genomic sequence. Studies of the regulation of two of the latter genes, termed sspG and sspJ, showed that both are expressed only in sporulation. The sspG gene is transcribed in the mother cell compartment by RNA polymerase with the mother cell-specific sigma factor for RNA polymerase, sigmaK, and is cotranscribed with a downstream gene, yurS; sspG transcription also requires the DNA binding protein GerE. In contrast, sspJ is transcribed in the forespore compartment by RNA polymerase with the forespore-specific sigmaG and appears to give a monocistronic transcript. A mutation eliminating SspG had no effect on sporulation or spore properties, while loss of SspJ caused a slight decrease in the rate of spore outgrowth in an otherwise wild-type background.
Figures


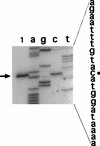
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

