Genetic and biochemical analyses of the tec operon suggest a route for evolution of chlorobenzene degradation genes
- PMID: 9864349
- PMCID: PMC103568
- DOI: 10.1128/JB.181.1.341-346.1999
Genetic and biochemical analyses of the tec operon suggest a route for evolution of chlorobenzene degradation genes
Abstract
The TecA broad-spectrum chlorobenzene dioxygenase of Burkholderia sp. strain PS12 catalyzes the first step in the mineralization of 1,2,4, 5-tetrachlorobenzene. The catabolic genes were localized on a small plasmid that belongs to the IncPbeta incompatibility group. PCR analysis of the genetic environment of the tec genes indicated high similarity to the transposon-organized catabolic tcb chlorobenzene degradation genes of Pseudomonas sp. strain P51. Sequence analysis of the regions flanking the tecA genes revealed an upstream open reading frame (ORF) with high similarity to the todF 2-hydroxy-6-oxo-2,4-heptadienoate hydrolase gene of Pseudomonas putida F1 and a discontinuous downstream ORF showing high similarity to the todE catechol 2,3-dioxygenase gene of strain F1. Both homologues in strain P51 exist only as deletion remnants. We suggest that different genetic events thus led to inactivation of the perturbing meta-cleavage enzymes in strains P51 and PS12 during the evolution of efficient chlorobenzene degradation pathways. Biochemical characterization of TodF-like protein TlpF and a genetically refunctionalized TodE-like protein, TlpE, produced in Escherichia coli provided data consistent with the proposed relationships.
Figures
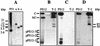


References
-
- Ahmad D, Fraser J, Sylvestre M, Larose A, Khan A, Bergeron J, Juteau J M, Sondossi M. Sequence of the bphD gene encoding 2-hydroxy-6-oxo-(phenyl/chlorophenyl)hexa-2,4-dienoic acid (HOP/cPDA) hydrolase involved in the biphenyl/polychlorinated biphenyl degradation pathway in Comamonas testosteroni: evidence suggesting involvement of Ser112 in catalytic activity. Gene. 1995;156:69–74. - PubMed
-
- Arand M, Grant D F, Beetham J K, Friedberg T, Oesch F, Hammock B D. Sequence similarity of mammalian epoxide hydrolases to the bacterial haloalkane dehalogenase and other related proteins. Implication for the potential catalytic mechanism of enzymatic epoxide hydrolysis. FEBS Lett. 1994;338:251–256. - PubMed
-
- Beil S, Happe B, Timmis K N, Pieper D H. Genetic and biochemical characterization of the broad spectrum chlorobenzene dioxygenase from Burkholderia sp. strain PS12: dechlorination of 1,2,4,5-tetrachlorobenzene. Eur J Biochem. 1997;247:190–199. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

